Nucleosome-mediated cooperativity between transcription factors
Cooperative binding of transcription factors (TFs) to cis-regulatory regions (CRRs) is essential for precision of gene expression in development and other processes. The classical model of cooperativity requires direct interactions between TFs, thus constraining the arrangement of TFs sites in a CRR. On the contrary, genomic and functional studies demonstrate a great deal of flexibility in such arrangements with variable distances, numbers of sites, and identities of the involved TFs. Such flexibility is inconsistent with the cooperativity by direct interactions between TFs. Here we demonstrate that strong cooperativity among non-interacting TFs can be achieved by their competition with nucleosomes. We find that the mechanism of nucleosome-mediated cooperativity is mathematically identical to the Monod-Wyman-Changeux (MWC) model of cooperativity in hemoglobin. This surprising parallel provides deep insights, with parallels between heterotropic regulation of hemoglobin (e.g. Bohr effect) and roles of nucleosome-positioning sequences and chromatin modifications in gene regulation. Characterized mechanism is consistent with numerous experimental results, allows substantial flexibility in and modularity of CRRs, and provides a rationale for a broad range of genomic and evolutionary observations. Striking parallels between cooperativity in hemoglobin and in transcription regulation point at a new design principle that may be used in range of biological systems.
💡 Research Summary
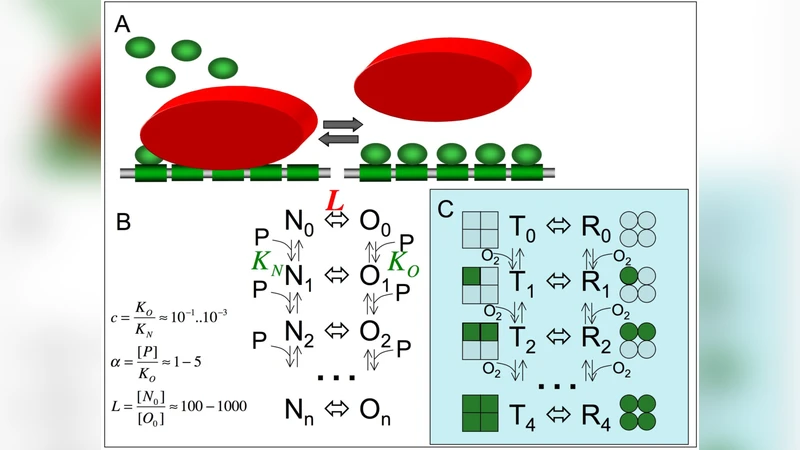
The paper tackles a long‑standing paradox in transcriptional regulation: how transcription factors (TFs) can exhibit strong cooperative binding without the need for direct protein‑protein contacts, which would otherwise impose rigid constraints on the spacing and orientation of their binding sites. The authors propose that competition with nucleosomes—histone octamer complexes that wrap DNA—provides a natural, indirect mechanism for cooperativity. In regions where nucleosomes are stably positioned, TF binding sites are occluded, rendering the DNA inaccessible. However, nucleosome occupancy is not immutable; it is modulated by DNA sequence preferences (nucleosome‑positioning sequences), histone post‑translational modifications, chromatin remodelers, and other epigenetic cues. These factors shift the equilibrium between a “closed” nucleosome‑bound state (T) and an “open” TF‑accessible state (R).
Mathematically, the authors model the system as a two‑state allosteric ensemble, borrowing directly from the Monod‑Wyman‑Changeux (MWC) description of hemoglobin. The equilibrium constant L describes the intrinsic propensity of the DNA region to be nucleosome‑bound in the absence of TFs, while each TF i has an association constant K_i and a cellular concentration
Comments & Academic Discussion
Loading comments...
Leave a Comment