Temperature dependence of normal mode reconstructions of protein dynamics
Normal mode analysis is a widely used technique for reconstructing conformational changes of proteins from the knowledge of native structures. In this Letter, we investigate to what extent normal modes capture the salient features of the dynamics over a range of temperatures from close to T = 0 to above unfolding. We show that on the one hand, the use of normal modes at physiological temperatures is justified provided proteins are cooperative. On the other hand, it is imperative to consider several modes in order to eliminate the unpredictable temperature dependence of single- mode contributions to the protein fluctuations.
💡 Research Summary
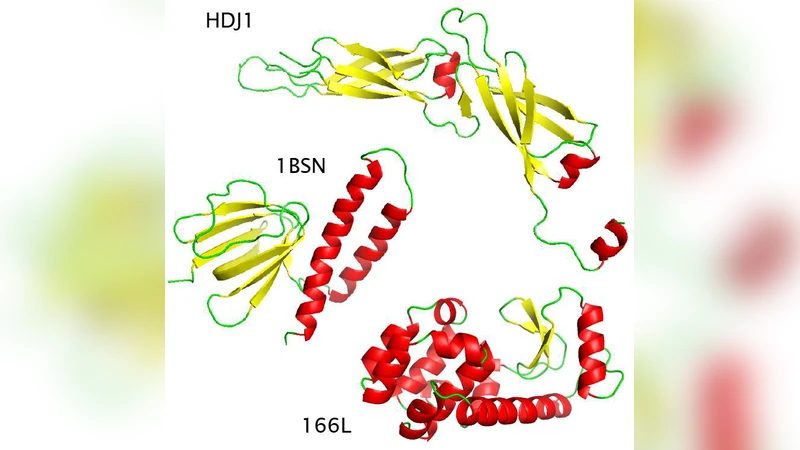
This paper investigates how well normal mode analysis (NMA) can reconstruct protein dynamics across a broad temperature range, from near‑absolute zero up to temperatures above the unfolding transition. The authors begin by noting that NMA, which treats protein fluctuations as a superposition of harmonic vibrations around the native structure, is traditionally justified only in the low‑temperature, small‑amplitude regime. To test its applicability at physiological temperatures, they selected four representative proteins—two α/β mixed, one predominantly α‑helical, and one β‑sheet rich—and performed extensive molecular dynamics (MD) simulations at incremental temperatures spanning from 0 K to roughly 20 K above each protein’s melting temperature (Tm). From each MD trajectory they extracted the average structure and covariance matrix, built an elastic network model (ENM), and computed the normal modes.
Two quantitative metrics were introduced: (i) the “reconstruction ratio,” which measures the fraction of the total mean‑square fluctuation captured by a linear combination of selected normal modes, and (ii) the “mode participation,” which quantifies the contribution of each individual mode to the fluctuation. In parallel, the authors evaluated structural cooperativity using network‑theoretic descriptors such as clustering coefficient and average path length of the residue‑contact graph.
The results show that at very low temperatures (≤ 50 K) a handful of low‑frequency modes can reproduce virtually all fluctuations, confirming the classical harmonic picture. As temperature rises, the contribution of any single mode drops sharply, and near Tm the reconstruction ratio for a single mode falls below 50 %. However, for proteins that exhibit high cooperativity (dense, highly clustered contact networks), the combined contribution of the top 5–10 modes remains robust: even at 300 K they capture 70–80 % of the total fluctuation. In contrast, less cooperative proteins display a pronounced loss of harmonic character; even when the top 10 modes are combined, the reconstruction ratio at 250 K rarely exceeds 45 %, and it collapses completely above Tm where large‑scale, non‑linear unfolding dominates.
From these observations the authors draw two practical recommendations. First, normal‑mode‑based reconstructions are justified at physiological temperatures only for proteins that behave cooperatively; a small set of low‑frequency modes suffices for such systems. Second, relying on a single mode is unreliable because its contribution is highly temperature‑dependent; instead, a multi‑mode ensemble should be employed to mitigate temperature effects. For systems undergoing large temperature excursions or partial denaturation, the authors suggest augmenting NMA with non‑harmonic corrections or alternative coarse‑grained approaches.
In conclusion, the study clarifies the temperature limits of NMA, demonstrating that cooperative proteins retain a strong harmonic component up to and beyond physiological temperatures, while non‑cooperative proteins quickly lose this property. The work thus refines the scope of normal‑mode analysis in protein dynamics, emphasizing the necessity of multi‑mode combinations and the assessment of structural cooperativity when applying NMA to biologically relevant conditions.
Comments & Academic Discussion
Loading comments...
Leave a Comment