Velocity oscillations in actin-based motility
We present a simple and generic theoretical description of actin-based motility, where polymerization of filaments maintains propulsion. The dynamics is driven by polymerization kinetics at the filaments’ free ends, crosslinking of the actin network, attachment and detachment of filaments to the obstacle interfaces and entropic forces. We show that spontaneous oscillations in the velocity emerge in a broad range of parameter values, and compare our findings with experiments.
💡 Research Summary
The paper presents a minimal yet comprehensive theoretical framework for actin‑driven motility, integrating filament polymerization dynamics, network cross‑linking elasticity, and the stochastic attachment–detachment processes that occur at the interface between actin filaments and a moving obstacle (e.g., a cell membrane or a pathogen surface). Polymerization at filament barbed ends is modeled as a force‑dependent rate proportional to the local G‑actin concentration, while depolymerization is accelerated by compressive forces acting on the filament tip. Cross‑linkers generate an effective elastic modulus and visco‑elastic relaxation time for the actin gel, providing a restoring force that opposes obstacle motion and stores mechanical energy. Attachment to the obstacle occurs with a basal on‑rate k_on, whereas detachment follows a force‑sensitive off‑rate k_off(F) that rises sharply once the compressive load exceeds a threshold, introducing a nonlinear feedback loop. Entropic forces arising from filament bending and confinement are also incorporated to capture steric repulsion near the obstacle.
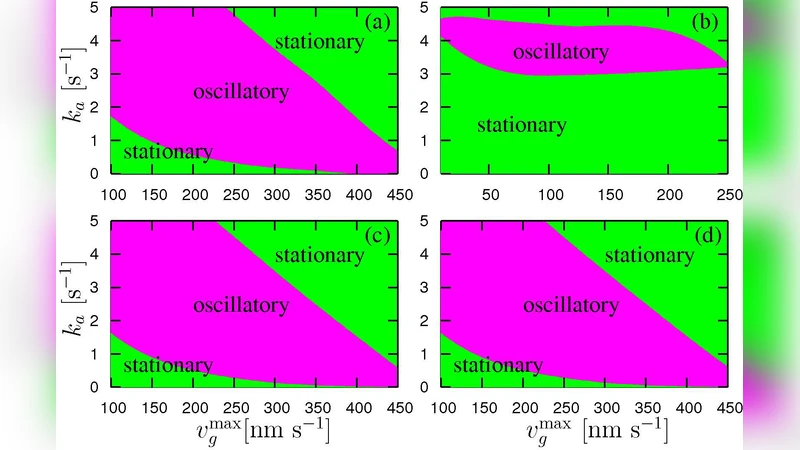
Mathematically the authors couple a one‑dimensional continuum equation for gel density and stress with a stochastic master equation governing the number of filaments in contact with the obstacle. The coupled system yields time‑dependent obstacle velocity V(t), filament contact number N(t), and gel stress σ(t). Numerical integration across a broad parameter space (polymerization rate k_p, baseline detachment rate k_off0, force‑sensitivity α, cross‑link stiffness G_c, and gel modulus E) reveals three distinct dynamical regimes. In the “steady‑propulsion” regime, high k_p and low α keep filaments attached, leading to a constant average velocity. In the “oscillatory” regime, intermediate values of k_p and α generate a delayed feedback: excessive attachment builds up gel stress, which then triggers a rapid increase in k_off(F), causing a sudden loss of contacts and a drop in velocity. The remaining filaments subsequently re‑polymerize, re‑attach, and the cycle repeats. This produces self‑sustained velocity oscillations with periods ranging from a few seconds to tens of seconds and amplitudes of 10–30 % of the mean speed. In the “suppressed‑motion” regime, weak cross‑linking or low gel stiffness prevents sufficient stress accumulation, and the system either stalls or moves at a very low, non‑oscillatory speed.
Parameter sweeps show that increasing k_p shortens the oscillation period and raises the amplitude up to a point; beyond a critical k_p the system locks into steady motion because detachment never becomes dominant. Raising the force‑sensitivity α amplifies both period and amplitude, as detachment becomes more abrupt. Strengthening cross‑linking (higher G_c or E) lengthens the period and damps the amplitude by storing more elastic energy and slowing the feedback. The authors validate the model against experimental data from Listeria monocytogenes and actin‑coated beads, reproducing observed average speeds (0.1–0.3 µm s⁻¹), oscillation periods (5–30 s), and speed fluctuations (≈10–30 % of the mean).
The key insight is that actin‑based propulsion is not governed solely by polymerization kinetics; rather, the interplay of nonlinear attachment–detachment dynamics and the visco‑elastic response of the cross‑linked network creates a delayed negative feedback that can naturally generate velocity oscillations. This unified description bridges molecular‑scale processes and mesoscopic motility behavior, offering a predictive tool for designing synthetic motile systems, interpreting cellular migration patterns, and engineering bio‑inspired nanomachines that exploit controlled oscillatory motion.
Comments & Academic Discussion
Loading comments...
Leave a Comment