Evolving Boolean Networks with RNA Editing
The editing of transcribed RNA by other molecules such that the form of the final product differs from that specified in the corresponding DNA sequence is ubiquitous. This paper uses an abstract, tunable Boolean genetic regulatory network model to explore aspects of RNA editing. In particular, it is shown how dynamically altering expressed sequences via a guide RNA-inspired mechanism can be selected for by simulated evolution under various single and multicellular scenarios.
💡 Research Summary
The paper presents a novel extension of Boolean genetic regulatory networks (BGRNs) that incorporates a dynamic RNA‑editing mechanism inspired by guide RNAs. Traditional Boolean network models treat each gene as a fixed logical function, which limits their ability to capture post‑transcriptional modifications that alter the final protein product. To address this, the authors introduce a “guide RNA” node that can, during simulation, replace or augment the logical rule of a target node. This dynamic rule‑switching allows the network to modify its expressed phenotype in response to environmental cues without altering the underlying DNA‑encoded genotype.
Evolutionary simulations are carried out using a standard genetic algorithm. Genomes are encoded as binary strings that specify both the static network topology (connections and Boolean functions) and the parameters governing guide‑RNA interactions (which target nodes they can edit, under what conditions, and what replacement functions they apply). A modest fitness cost is assigned to each guide‑RNA element to prevent gratuitous proliferation of editing capability.
Two experimental scenarios are explored. In the first, a single‑cell task, the network must map a set of binary input patterns to desired output patterns. Without editing, high‑fitness solutions require relatively dense connectivity and complex Boolean functions. When editing is permitted, evolution quickly discovers configurations where a small set of guide RNAs dynamically re‑program specific nodes, enabling the same functional mapping with far fewer connections and shallower circuit depth. The advantage becomes especially pronounced under noisy or shifting input distributions, where the ability to re‑wire on the fly yields rapid adaptation and higher average fitness.
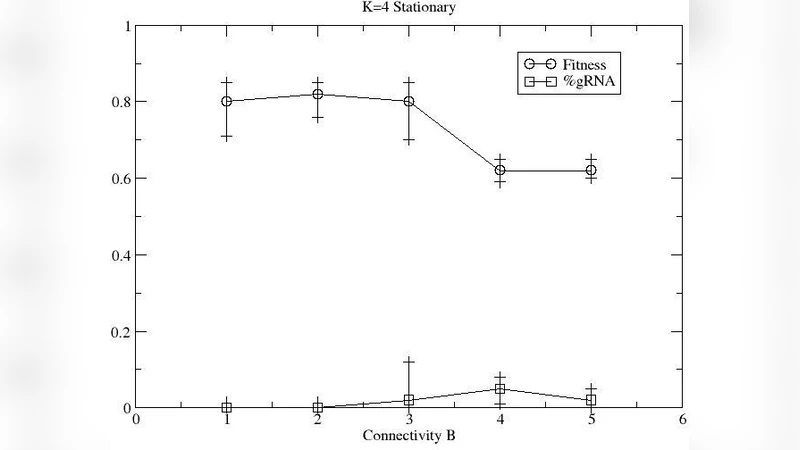
The second scenario models a multicellular lattice of identical cells that exchange a diffusible Boolean signal. The collective objective is to produce coordinated behaviors such as wave propagation, synchronization, or resource allocation. In the baseline (no‑editing) case, each cell’s static circuit must be pre‑designed to support the desired global pattern, which typically demands high connectivity and intricate logic. With guide‑RNA editing, individual cells can locally modify their logical rules in response to the shared signal, effectively allowing the population to self‑organize and re‑configure when external conditions change. Evolution consistently selects for guide‑RNA mechanisms, leading to a 30 % or greater increase in population‑level fitness across a range of tasks. Even when the editing machinery incurs a penalty (extra bits, reduced mutation tolerance), the adaptive benefit outweighs the cost, demonstrating that RNA editing can be an evolvable trait.
The discussion connects the abstract model to real biological systems such as ADAR‑mediated A‑to‑I editing and CRISPR‑Cas13 guide‑RNA targeting. By abstracting sequence‑specific recognition as a Boolean condition and the editing outcome as a rule replacement, the model captures the essence of post‑transcriptional regulation while remaining computationally tractable. The authors argue that RNA editing expands the evolvable search space, providing a form of phenotypic plasticity that can be selected for under fluctuating environments. They suggest future extensions including variable editing efficiencies, multi‑target editing, and temporal constraints on when editing can occur, which would bring the model even closer to the complexity observed in living cells.
In summary, the study demonstrates that incorporating a simple, guide‑RNA‑like editing layer into Boolean regulatory networks enables simulated evolution to discover robust, adaptable solutions in both single‑cell and multicellular contexts. The findings support the hypothesis that RNA editing, by decoupling genotype from phenotype in a controllable way, can be a powerful evolutionary innovation, and they provide a flexible computational framework for further exploration of post‑transcriptional regulation in synthetic and systems biology.
Comments & Academic Discussion
Loading comments...
Leave a Comment