On Joint Diagonalisation for Dynamic Network Analysis
Joint diagonalisation (JD) is a technique used to estimate an average eigenspace of a set of matrices. Whilst it has been used successfully in many areas to track the evolution of systems via their eigenvectors; its application in network analysis is novel. The key focus in this paper is the use of JD on matrices of spanning trees of a network. This is especially useful in the case of real-world contact networks in which a single underlying static graph does not exist. The average eigenspace may be used to construct a graph which represents the `average spanning tree’ of the network or a representation of the most common propagation paths. We then examine the distribution of deviations from the average and find that this distribution in real-world contact networks is multi-modal; thus indicating several \emph{modes} in the underlying network. These modes are identified and are found to correspond to particular times. Thus JD may be used to decompose the behaviour, in time, of contact networks and produce average static graphs for each time. This may be viewed as a mixture between a dynamic and static graph approach to contact network analysis.
💡 Research Summary
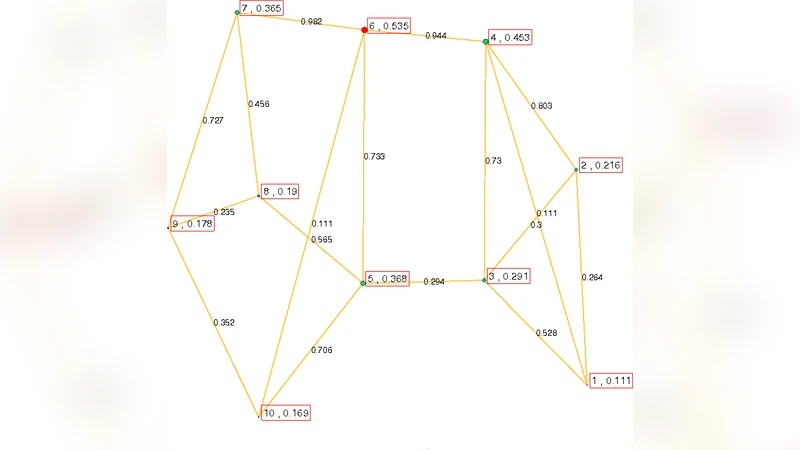
The paper introduces a novel application of Joint Diagonalisation (JD) to dynamic contact networks by operating on the adjacency matrices of spanning trees generated from the network. Traditional static‑graph approaches collapse temporal information and can create spurious links, whereas the proposed method first samples a large number of spanning trees via breadth‑first searches from randomly chosen root nodes. Each sampled tree Hᵢ is expressed as Hᵢ = U Cᵢ Uᵀ, where U is a common orthogonal matrix and Cᵢ is diagonal. JD finds the orthogonal matrix \bar{U} that minimises the sum of squared off‑diagonal elements across all Cᵢ, effectively yielding an average eigenspace. The average sampling matrix \bar{H}= \bar{U},\bar{C},\bar{U}ᵀ then encodes, in its weighted edges, the proportion of spanning trees that use each link – a representation the authors call the “average spanning tree”.
The authors evaluate the method on three real Bluetooth contact datasets (Cambridge, Infocom06, MIT) and on a synthetic network with known structure. For each dataset they generate 10 000 spanning trees, apply JD, and compute the deviation δᵢ of each tree from \bar{H}. Kernel‑smoothed histograms of δᵢ reveal multimodal distributions, indicating that the underlying contact process operates in distinct temporal modes (e.g., class‑time, night‑time, break periods). Gaussian mixture modelling isolates these modes, and mode‑specific average graphs \bar{H} expose characteristic structures such as bridges formed by particular nodes (e.g., nodes 3 and 20 in the Cambridge data) that dominate during specific periods.
To demonstrate practical relevance, the authors run SIR epidemic simulations seeded at nodes identified as bridges. The simulations confirm that infections initiated in the identified modes spread more rapidly, validating the predictive power of the JD‑derived graphs. Additional experiments show that deliberately biasing the sampling (removing a high‑frequency link) leads JD to down‑weight that link in \bar{H}, illustrating the method’s sensitivity to underlying propagation pathways.
Overall, the study shows that JD can compress a temporally rich contact network into a set of static, mode‑specific graphs while preserving essential dynamical information. This hybrid static‑dynamic representation enables efficient analysis of routing, epidemic spreading, and other processes on time‑varying networks, offering a powerful new tool for researchers dealing with complex, temporally evolving graph data.
Comments & Academic Discussion
Loading comments...
Leave a Comment