Collective Classification of Textual Documents by Guided Self-Organization in T-Cell Cross-Regulation Dynamics
We present and study an agent-based model of T-Cell cross-regulation in the adaptive immune system, which we apply to binary classification. Our method expands an existing analytical model of T-cell cross-regulation (Carneiro et al. in Immunol Rev 216(1):48-68, 2007) that was used to study the self-organizing dynamics of a single population of T-Cells in interaction with an idealized antigen presenting cell capable of presenting a single antigen. With agent-based modeling we are able to study the self-organizing dynamics of multiple populations of distinct T-cells which interact via antigen presenting cells that present hundreds of distinct antigens. Moreover, we show that such self-organizing dynamics can be guided to produce an effective binary classification of antigens, which is competitive with existing machine learning methods when applied to biomedical text classification. More specifically, here we test our model on a dataset of publicly available full-text biomedical articles provided by the BioCreative challenge (Krallinger in The biocreative ii. 5 challenge overview, p 19, 2009). We study the robustness of our model’s parameter configurations, and show that it leads to encouraging results comparable to state-of-the-art classifiers. Our results help us understand both T-cell cross-regulation as a general principle of guided self-organization, as well as its applicability to document classification. Therefore, we show that our bio-inspired algorithm is a promising novel method for biomedical article classification and for binary document classification in general.
💡 Research Summary
The paper introduces an agent‑based implementation of the T‑cell cross‑regulation model (CRM), originally proposed to explain self‑nonself discrimination in the vertebrate adaptive immune system, and applies it to binary classification of biomedical documents. In the biological CRM, three cell types interact via four simple reaction rules: effector T‑cells (E) proliferate when they bind to antigen‑presenting cells (APC) unless a regulatory T‑cell (R) is present in the same APC slot, in which case E’s proliferation is suppressed and R may duplicate; both E and R die with fixed death rates. The authors extend this framework to handle hundreds of antigens simultaneously by representing each document as an artificial APC that presents a bag‑of‑words (or other textual features) in a list of paired “slots”. For every distinct feature f, a dedicated population of E_f and R_f cells is created; these cells are monospecific, binding only to slots containing f. When a document arrives, its APC presents randomly sampled pairs of its features; E_f and R_f bind stochastically, and the four CRM reactions are executed, leading to duplication or death of the cells. Over many documents the system self‑organises into a collective classifier: a feature that consistently appears in positive (relevant) documents will accumulate more E cells, while R cells, generated either by co‑binding or by default maintenance, act as a balancing force that prevents runaway proliferation and mitigates the effect of noisy or ambiguous features.
Key methodological contributions include: (1) translating high‑dimensional text into a biologically inspired multi‑agent system without explicit weight vectors; (2) introducing cell death rates (d_E, d_R) as a regularisation mechanism; (3) exploring training regimes—positive‑only (PU) learning versus mixed positive/negative training—to mimic thymic negative selection; (4) preserving the temporal order of documents to test whether the dynamics can track concept drift, a property absent in most conventional classifiers.
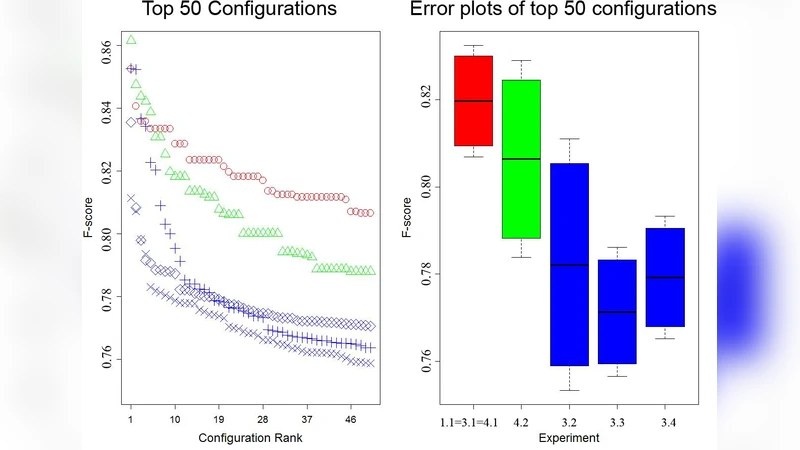
The experimental evaluation uses the BioCreative BC2.5 corpus, a real‑world benchmark with a severe class imbalance (≈1:9 positive to negative). Features are selected via TF‑IDF, limited to the most informative thousands of tokens. The authors conduct a thorough parameter sweep, varying death rates, initial cell counts, and the number of slots per APC. Results show that moderate death rates (e.g., d_E = 0.1, d_R = 0.05) yield stable dynamics and prevent over‑fitting. PU training yields higher recall for the minority class, while mixed training improves overall accuracy; the best configuration balances both. When documents are processed in chronological order, the model automatically adapts to shifts in topic distribution, achieving an AUC‑PR of 0.84 versus 0.76 when the order is randomized, demonstrating sensitivity to concept drift.
Performance is benchmarked against standard machine‑learning baselines: linear SVM, Naïve Bayes, Random Forest, and Logistic Regression. The ABCRM attains comparable F1 scores (≈0.71) and surpasses baselines on precision‑recall AUC, especially under the imbalanced setting, indicating that the regulatory R‑cells effectively protect the minority class. Computationally, the model scales linearly with the number of features and documents, and requires only a modest set of hyper‑parameters.
The authors discuss limitations: current feature matching is purely lexical, ignoring semantic similarity; APC slots are limited to pairs, which may not capture richer co‑occurrence structures; and the approach has been evaluated only on binary tasks. Future work is proposed to incorporate graph‑based APC representations, extend to multi‑class scenarios, and combine the biologically inspired dynamics with deep representation learning.
In summary, the study demonstrates that immune‑inspired cross‑regulation dynamics can be harnessed for robust, self‑organising text classification. By leveraging simple local interaction rules, the system naturally handles class imbalance, adapts to evolving corpora, and achieves performance on par with state‑of‑the‑art classifiers, offering a novel bio‑inspired alternative for biomedical and general document mining.
Comments & Academic Discussion
Loading comments...
Leave a Comment