Statistical methods for tissue array images - algorithmic scoring and co-training
Recent advances in tissue microarray technology have allowed immunohistochemistry to become a powerful medium-to-high throughput analysis tool, particularly for the validation of diagnostic and prognostic biomarkers. However, as study size grows, the manual evaluation of these assays becomes a prohibitive limitation; it vastly reduces throughput and greatly increases variability and expense. We propose an algorithm - Tissue Array Co-Occurrence Matrix Analysis (TACOMA) - for quantifying cellular phenotypes based on textural regularity summarized by local inter-pixel relationships. The algorithm can be easily trained for any staining pattern, is absent of sensitive tuning parameters and has the ability to report salient pixels in an image that contribute to its score. Pathologists’ input via informative training patches is an important aspect of the algorithm that allows the training for any specific marker or cell type. With co-training, the error rate of TACOMA can be reduced substantially for a very small training sample (e.g., with size 30). We give theoretical insights into the success of co-training via thinning of the feature set in a high-dimensional setting when there is “sufficient” redundancy among the features. TACOMA is flexible, transparent and provides a scoring process that can be evaluated with clarity and confidence. In a study based on an estrogen receptor (ER) marker, we show that TACOMA is comparable to, or outperforms, pathologists’ performance in terms of accuracy and repeatability.
💡 Research Summary
The paper introduces TACOMA (Tissue Array Co‑Occurrence Matrix Analysis), a novel algorithm for automated scoring of tissue microarray (TMA) images that combines gray‑level co‑occurrence matrices (GLCM) with Random Forest (RF) classification. Traditional TMA scoring tools rely heavily on intensity thresholds, color filters, segmentation, and extensive parameter tuning, which limits their robustness and scalability. TACOMA avoids these pitfalls by using only the spatial relationships of pixel intensities to capture texture information, thereby eliminating the need for complex preprocessing.
A key innovation is the use of a small set of expert‑selected image patches to create a feature mask. The GLCM is computed for each image (or patch) across predefined spatial relationships; only the entries that correspond to the mask—those that appear frequently in the expert patches—are retained as features. This implicit, non‑parametric feature selection embeds domain knowledge while requiring minimal manual effort and prevents background or non‑specific tissue from influencing the classifier.
All retained GLCM entries (often thousands) are fed directly into a Random Forest classifier. RF’s ensemble of decision trees, each built on a random subset of features, naturally handles the high‑dimensional texture space, captures nonlinear interactions, and provides built‑in variable importance for interpretability. The authors demonstrate that RF outperforms support vector machines, boosting, and naive Bayes on the same data, achieving an accuracy of 78.57% compared with 65.24% for SVM and 61.28% for boosting.
To address the common problem of limited labeled data in biomedical studies, the authors incorporate co‑training. They split the high‑dimensional feature set into two “views” (by random thinning) and train two independent RF models. Each model iteratively adds its most confident predictions to the training set of the other, effectively enlarging the labeled pool without additional expert annotation. The paper provides theoretical justification for this thinning approach, showing that when the two views are sufficiently redundant, the classification power of each thinned subset matches that of the full feature set. Empirically, with as few as 30 initially labeled samples, co‑training reduces error rates substantially.
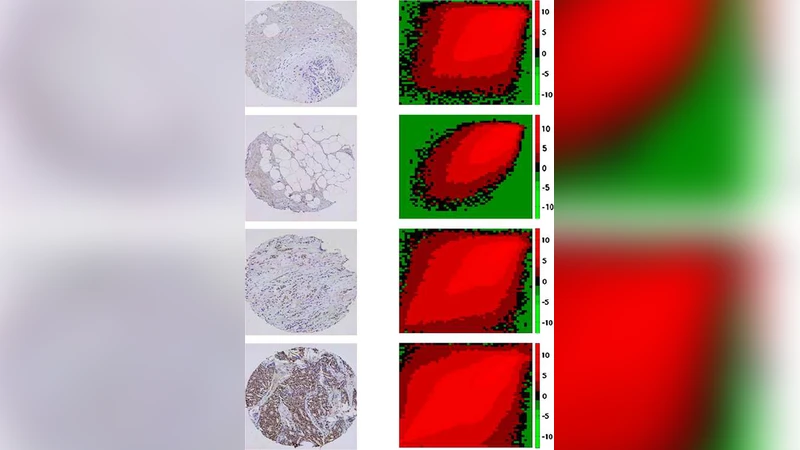
The method is evaluated on a real‑world dataset of estrogen receptor (ER) stained TMA slides, scored on a four‑point scale (0–3). TACOMA’s performance is comparable to, and in some cases exceeds, that of experienced pathologists in both accuracy and repeatability. An additional benefit is the generation of saliency maps that highlight the pixels contributing most to the final score, enhancing interpretability and allowing pathologists to verify algorithmic decisions.
Strengths of the work include (1) a parameter‑free texture representation that is robust across staining patterns, (2) a lightweight expert‑guided feature mask that integrates domain knowledge without manual feature engineering, (3) an effective co‑training scheme that mitigates the scarcity of labeled data, and (4) transparent output through pixel‑level contribution visualizations. Limitations involve the need to pre‑define the number of gray levels (N_g) for GLCM computation, potential sensitivity to image quality, and the fact that the current validation focuses on a single marker (ER) and a single institution’s data. Future directions suggested by the authors are extending the approach to multiple markers and subcellular compartments, exploring hybrid models that combine deep‑learning texture extractors with the GLCM‑RF pipeline, and conducting multi‑center studies to assess generalizability. Overall, TACOMA represents a significant step toward scalable, reproducible, and interpretable automated scoring of high‑throughput tissue microarray experiments.
Comments & Academic Discussion
Loading comments...
Leave a Comment