Data Sharing Options for Scientific Workflows on Amazon EC2
Efficient data management is a key component in achieving good performance for scientific workflows in distributed environments. Workflow applications typically communicate data between tasks using files. When tasks are distributed, these files are either transferred from one computational node to another, or accessed through a shared storage system. In grids and clusters, workflow data is often stored on network and parallel file systems. In this paper we investigate some of the ways in which data can be managed for workflows in the cloud. We ran experiments using three typical workflow applications on Amazon’s EC2. We discuss the various storage and file systems we used, describe the issues and problems we encountered deploying them on EC2, and analyze the resulting performance and cost of the workflows.
💡 Research Summary
The paper investigates how scientific workflow data can be managed efficiently on Amazon EC2, where the choice of storage directly influences both execution time and monetary cost. Three representative workflows—Montage (an I/O‑intensive astronomy image‑mosaicking pipeline), Epigenomics (a memory‑intensive bio‑informatics pipeline), and CyberShake (a CPU‑intensive seismic‑simulation pipeline)—are used as testbeds because they span a wide range of data‑access patterns.
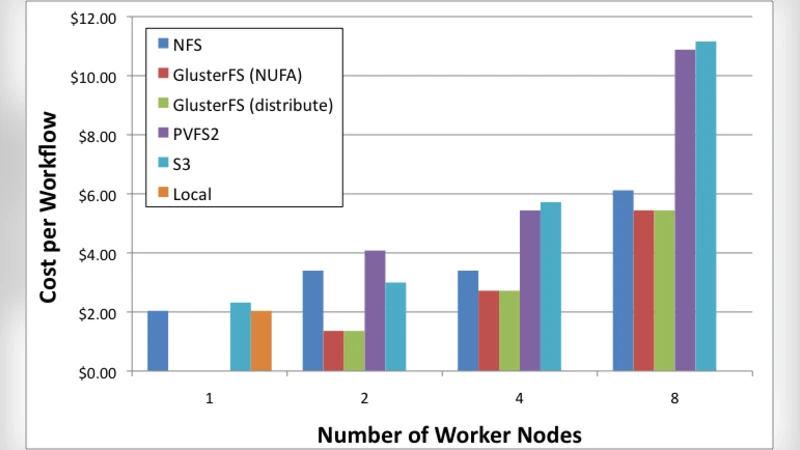
The authors evaluate five storage configurations that are realistic in a cloud setting: (1) per‑instance Amazon Elastic Block Store (EBS) volumes used as local disks, (2) shared EBS volumes mounted simultaneously on multiple instances, (3) Amazon Simple Storage Service (S3) as an object store, (4) a traditional Network File System (NFS) server deployed on a dedicated EC2 instance, and (5) a distributed file system (GlusterFS) spread across several instances. Each configuration is examined for its impact on (a) workflow makespan, (b) data‑transfer volume, (c) network utilization, and (d) total cost (including compute, storage, and data‑transfer charges).
Results show that the I/O‑heavy Montage workflow benefits most from the low‑latency, high‑throughput characteristics of local EBS disks; however, as the number of parallel tasks grows, the need to replicate intermediate files across instances inflates storage cost and reduces overall cost‑effectiveness. In contrast, the Epigenomics and CyberShake pipelines, which involve substantial intermediate data reuse, achieve better performance when a shared POSIX‑compatible file system is used. GlusterFS, in particular, provides automatic replication and self‑healing, which reduces network traffic and improves fault tolerance, albeit at the expense of higher management overhead and a modest performance penalty due to metadata synchronization.
S3 emerges as the most economical option for storing transient data or final results because of its pay‑as‑you‑go pricing and virtually unlimited durability. Nevertheless, the object‑store’s per‑request latency and the overhead of uploading/downloading many small files can dominate the runtime of workflows that generate a large number of short‑lived files. A hybrid approach—using S3 for bulk, infrequently accessed data while keeping hot intermediate files on a shared file system—delivers the best cost‑performance balance for most scenarios.
The paper also highlights a subtle but critical issue: EC2’s virtualization layer can break POSIX cache consistency when the same EBS volume is mounted by multiple instances. Inconsistent caches may lead to stale reads or lost writes. The authors mitigate this risk by enforcing file‑level locking, issuing periodic sync calls, and, when possible, avoiding shared‑volume configurations altogether.
Building on these observations, the authors propose a policy‑driven framework that integrates workflow scheduling with storage selection. The framework classifies each task according to its input/output size, expected compute duration, and user‑specified budget, then automatically maps the task to the most suitable storage backend (local EBS, shared EBS, NFS, GlusterFS, or S3). This dynamic mapping reduces the need for manual tuning and enables the system to adapt to changing workload characteristics or pricing fluctuations.
In summary, the study demonstrates that no single storage solution dominates across all scientific workflows on EC2. Instead, a mixed‑storage strategy—tailored to the specific I/O profile of each task and balanced against cost constraints—yields the most efficient execution. The findings provide concrete guidance for researchers and cloud architects seeking to deploy data‑intensive scientific pipelines in public‑cloud environments, and they lay the groundwork for automated, cost‑aware workflow management systems.
Comments & Academic Discussion
Loading comments...
Leave a Comment