Machine learning of hierarchical clustering to segment 2D and 3D images
We aim to improve segmentation through the use of machine learning tools during region agglomeration. We propose an active learning approach for performing hierarchical agglomerative segmentation from superpixels. Our method combines multiple features at all scales of the agglomerative process, works for data with an arbitrary number of dimensions, and scales to very large datasets. We advocate the use of variation of information to measure segmentation accuracy, particularly in 3D electron microscopy (EM) images of neural tissue, and using this metric demonstrate an improvement over competing algorithms in EM and natural images.
💡 Research Summary
The paper presents a novel framework for image segmentation that integrates machine‑learning‑driven decision making into the hierarchical agglomeration stage, addressing both 2‑D natural images and 3‑D electron‑microscopy (EM) volumes. Traditional pipelines first over‑segment an image into superpixels (or supervoxels) and then merge these regions using simple heuristics such as the mean boundary probability. While effective for small regions, these heuristics degrade as regions grow, especially in EM data where thin neuronal processes and intricate topologies make even tiny boundary errors catastrophic for downstream connectomics.
To overcome these limitations, the authors propose an active‑learning approach that continuously generates training examples during the agglomeration itself. Starting from an initial over‑segmentation S, a Region Adjacency Graph (RAG) is built where each node corresponds to a region and edges connect adjacent regions. For every edge (u, v) a feature vector f(G, u, v) is computed. Features are drawn from two levels: (1) pixel‑level cues such as boundary probability maps, texton histograms, quantiles, moments, and Jensen‑Shannon divergence; (2) mid‑level cues that capture region orientation (derived from second‑moment matrices) and convex‑hull geometry (individual and combined volumes, ratios). These multi‑scale descriptors provide rich information across the entire hierarchy, from the smallest superpixels to large merged objects.
The merge priority function (MPF) π is modeled as a composition of a feature map f and a binary classifier c (π = c ∘ f). The classifier predicts a probability that two adjacent regions should be merged. The active‑learning loop works as follows: an initial policy (often the mean boundary probability or a random ranking) is used to propose the next merge. The proposed merge is compared against a gold‑standard segmentation A* derived from manual annotation. If the merge is correct (both regions belong to the same ground‑truth object) the edge is labeled “‑1” (merge) and the regions are actually merged; otherwise it receives label “+1” (do not merge) and the merge is rejected. The (feature, label) pair is added to the training set, and the classifier is retrained after each epoch. By repeating this process until the algorithm’s segmentation matches A*, the policy learns from examples spanning every scale of the hierarchy, eliminating the “feature‑space holes” that plague static training sets.
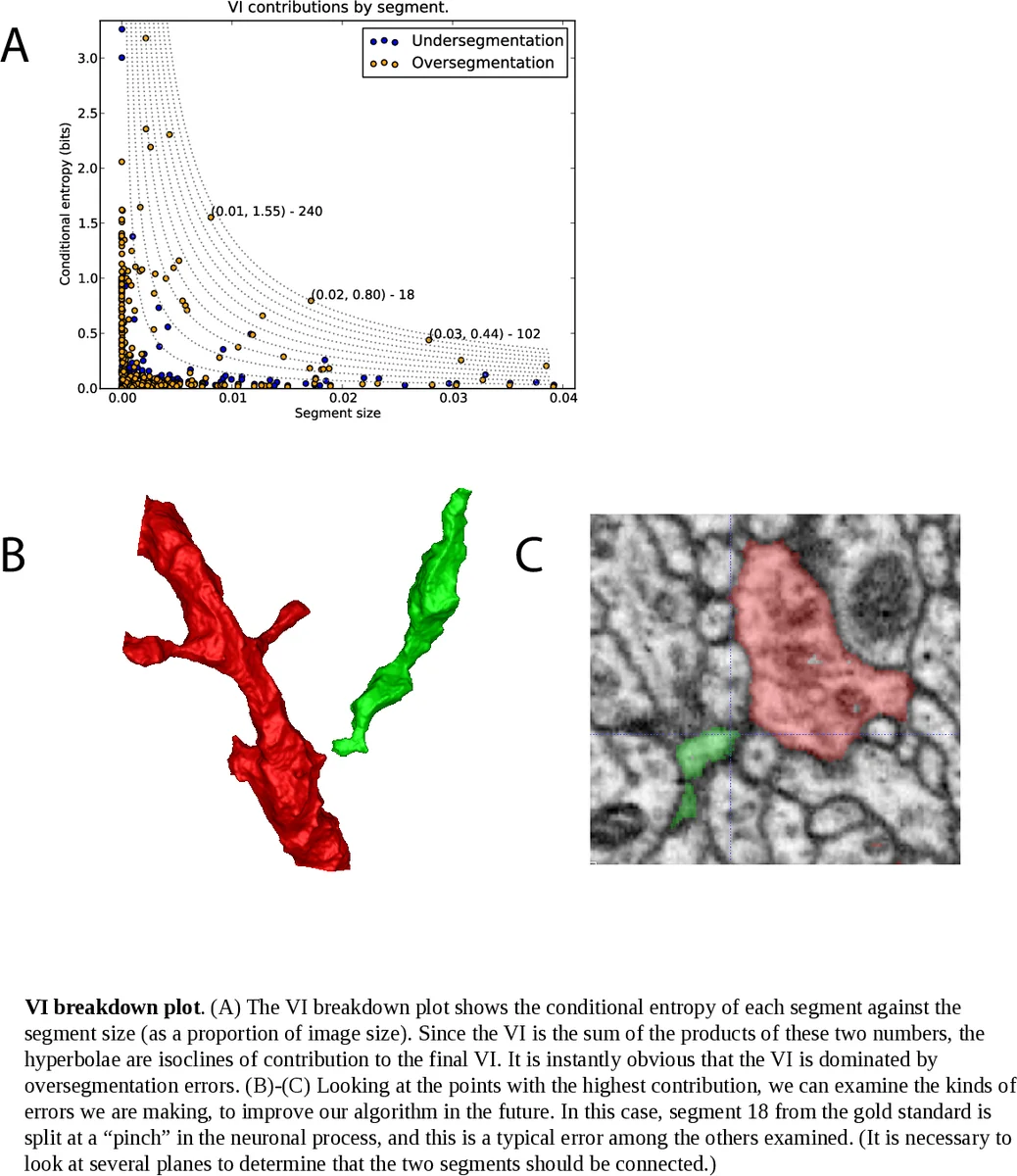
For evaluation the authors argue that region‑based metrics are more appropriate than boundary‑based precision‑recall, especially for EM where topological correctness outweighs pixel‑wise boundary accuracy. They adopt Variation of Information (VI), defined as the sum of conditional entropies H(S|U) + H(U|S) between a candidate segmentation S and ground truth U. VI penalizes both over‑segmentation (splitting true objects) and under‑segmentation (merging distinct objects) and is sensitive to topological errors.
Experiments on a large EM dataset (500 × 500 × 500 voxels, subdivided into 8 sub‑volumes) and on standard natural‑image benchmarks (e.g., BSDS500) demonstrate that the learned agglomeration consistently yields lower VI scores than baseline methods that rely solely on mean boundary probability, as well as improvements over previously published learning‑based agglomeration approaches such as LASH. In natural images the method also achieves higher boundary precision‑recall F‑measures, confirming its generality beyond the specialized EM domain.
All components—superpixel generation, RAG construction, feature extraction, and the active‑learning agglomeration—are implemented in an open‑source Python library called Gala (Graph‑based Active Learning of Agglomeration). Gala is dimension‑agnostic, supporting 2‑D, 3‑D, and even 4‑D (spatiotemporal) data with a unified API, enabling researchers to apply the technique to massive connectomics volumes or any other multi‑dimensional imaging problem.
In summary, the paper introduces a principled, scalable, and data‑driven method for hierarchical image segmentation. By actively learning merge decisions across the full hierarchy and leveraging rich multi‑scale features, the approach outperforms traditional heuristics and earlier learning‑based methods, particularly in challenging 3‑D EM datasets where accurate topology is essential. The open‑source Gala framework further amplifies the impact by making the technique readily accessible to the broader computer‑vision and neuroscience communities.
Comments & Academic Discussion
Loading comments...
Leave a Comment