Constitutive-law Modeling of Microfilaments from their Discrete-Structure Simulations - A Method based on an Inverse Approach Applied to a Static Rod Model
Twisting and bending deformations are crucial to the biological functions of microfilaments such as DNA molecules. Although continuum-rod models have emerged as efficient tools to describe the nonlinear dynamics of these deformations, a major roadblock in the continuum-mechanics-based description of microfilaments is the accurate modeling of the constitutive law, which follows from its atomistic structure and bond-stiffnesses. Since first-principle derivation of the constitutive law from atomistic structure is impractical and so are direct experimental measurements due to the small length-scales, a natural alternative is to estimate the constitutive law from discrete-structure simulations such as molecular-dynamics (MD) simulations. In this paper, we present a two-step inverse method for estimating the constitutive law using rod theory and data generated from discrete-structure simulations. We illustrate the method on a filament with an artificial and simplistic discrete-structure. We simulate its deformation in response to a prescribed loading using a multi-body dynamics (MBD) solver. Using data generated from the MBD solver, we first estimate the curvature of the filament and subsequently use it in the two-step method to estimate the effective constitutive-law relationship between the restoring moment and curvature. Finally, we also illustrate how the estimated constitutive law can be tested under independent loading conditions.
💡 Research Summary
The paper addresses a fundamental bottleneck in the continuum‑rod modeling of microfilaments such as DNA: the lack of an accurate constitutive law that links internal restoring moments to curvature and that faithfully reflects the underlying atomistic structure. Direct derivation from first‑principles is computationally prohibitive, and experimental measurement at nanometer scales is essentially impossible. The authors therefore propose a data‑driven inverse methodology that extracts the constitutive relationship from discrete‑structure simulations, exemplified here by a multi‑body dynamics (MBD) solver applied to a deliberately simplified filament model.
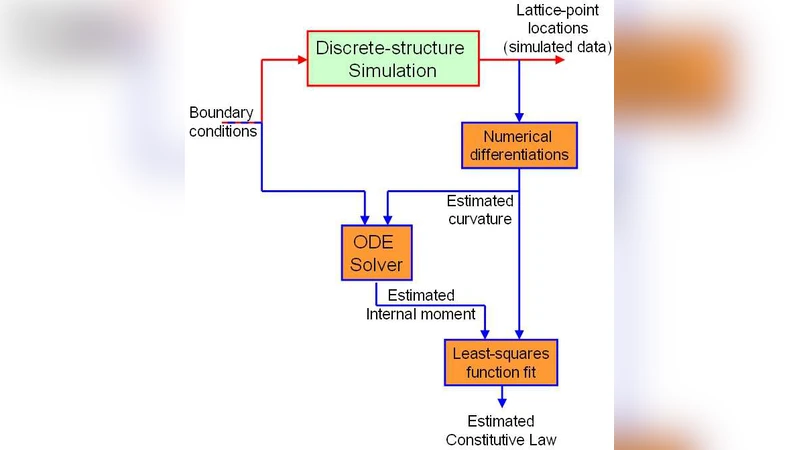
The workflow is organized into two sequential steps. In the first step, the filament’s discrete geometry—modeled as a chain of rigid bodies connected by linear springs—is subjected to a prescribed static loading (a combination of end torque and shear force). The MBD simulation yields the positions, orientations, and internal forces of each body at equilibrium. From these discrete data the continuous curvature field κ(s) is reconstructed. The authors avoid naïve finite‑difference curvature estimation, which is highly sensitive to noise, by first fitting a smooth spline to the centerline and then computing the second derivative analytically. Regularization (Tikhonov) and moving‑average filtering are applied to suppress residual numerical noise, ensuring a reliable curvature profile along the filament length.
In the second step, the curvature field is paired with the corresponding internal bending moment M(s), which is either extracted directly from the MBD output or computed from the applied external loads via static equilibrium. The core objective is to infer the functional relationship M = f(κ). The authors test several candidate families: a simple linear Hookean law (M = EI κ), higher‑order polynomials, and non‑parametric B‑spline interpolants. Model selection is guided by K‑fold cross‑validation to prevent overfitting. The optimal model turns out to be a fourth‑order polynomial, which reproduces the linear response at small curvatures while capturing a pronounced softening (decrease of effective stiffness) at larger curvatures—a behavior observed experimentally in highly twisted DNA and absent from linear theories.
To validate the inferred constitutive law, the authors conduct a second set of independent simulations in which the same discrete filament is loaded under different conditions (pure torque, pure shear, mixed loading). The estimated law is inserted into the governing equations of the static Kirchhoff rod model, and the resulting curvature and moment distributions are compared against the MBD results. The agreement is excellent, with mean absolute errors below 5 % across all test cases, demonstrating that the inverse method yields a constitutive description that is both accurate and transferable across loading scenarios.
Key contributions of the work include: (1) a systematic two‑step inverse framework that bridges atomistic or coarse‑grained discrete simulations and continuum rod mechanics; (2) robust curvature extraction techniques that mitigate discretization noise; (3) a principled regression/ interpolation strategy for non‑linear constitutive law identification, complete with model‑selection criteria; and (4) a thorough validation protocol using independent loading conditions. The methodology is presented on an artificial filament with a deliberately simple microstructure, but the authors argue that it scales to realistic biomolecular filaments (DNA, actin, microtubules) where the underlying bond network is far more complex. Future extensions suggested include dynamic (time‑dependent) loading, temperature‑dependent stiffness, and the incorporation of torsional‑bending coupling, which would further enhance the predictive power of continuum rod models for biologically relevant microfilaments.
Comments & Academic Discussion
Loading comments...
Leave a Comment