The Function of Communities in Protein Interaction Networks at Multiple Scales
Background: If biology is modular then clusters, or communities, of proteins derived using only protein interaction network structure should define protein modules with similar biological roles. We investigate the link between biological modules and network communities in yeast and its relationship to the scale at which we probe the network. Results: Our results demonstrate that the functional homogeneity of communities depends on the scale selected, and that almost all proteins lie in a functionally homogeneous community at some scale. We judge functional homogeneity using a novel test and three independent characterizations of protein function, and find a high degree of overlap between these measures. We show that a high mean clustering coefficient of a community can be used to identify those that are functionally homogeneous. By tracing the community membership of a protein through multiple scales we demonstrate how our approach could be useful to biologists focusing on a particular protein. Conclusions: We show that there is no one scale of interest in the community structure of the yeast protein interaction network, but we can identify the range of resolution parameters that yield the most functionally coherent communities, and predict which communities are most likely to be functionally homogeneous.
💡 Research Summary
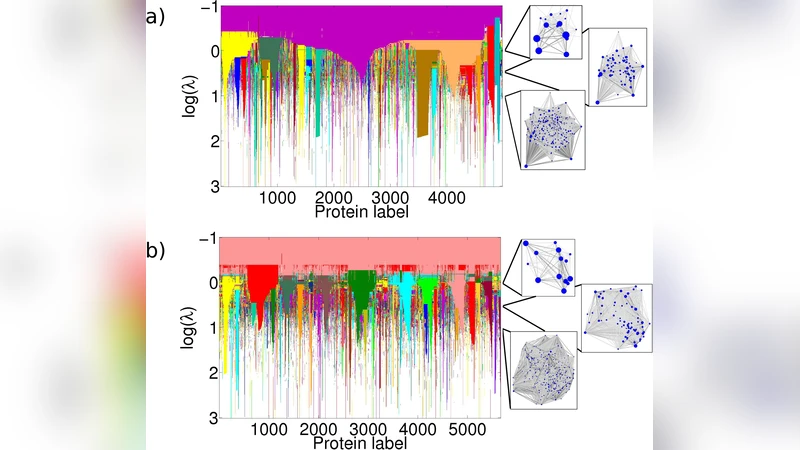
The paper investigates whether communities (clusters) identified solely from the topology of a protein‑protein interaction (PPI) network correspond to biologically meaningful modules, and how this correspondence depends on the scale at which the network is examined. Using two yeast PPI datasets—a high‑confidence “core” network and a more inclusive “full” network—the authors apply the Louvain community‑detection algorithm while varying its resolution parameter γ from 0.5 to 5.0. This generates a series of partitions ranging from a few large communities (low γ) to many small ones (high γ).
To assess functional homogeneity, three independent characterizations of protein function are employed: (1) Gene Ontology (GO) Biological Process term overlap, (2) MIPS functional category overlap, and (3) co‑expression similarity derived from a large microarray compendium. For each community and each functional measure, the authors compute an average similarity score and compare it against a null distribution generated by repeatedly sampling random protein sets of the same size. Communities with a Z‑score > 2 (p < 0.01) for a given measure are deemed functionally homogeneous.
The results reveal a strong dependence of functional homogeneity on the resolution parameter. Communities obtained at intermediate γ values (approximately 1.5–2.5) exhibit the highest proportion of statistically significant functional coherence across all three measures. In contrast, low‑resolution partitions produce overly large, heterogeneous modules, while high‑resolution partitions become too fragmented to achieve statistical power. Importantly, the authors discover that the mean clustering coefficient (C) of a community is a powerful predictor of functional homogeneity: communities with C ≥ 0.4 are enriched for significant GO, MIPS, and expression coherence (≈ 85 % of such communities pass the test). This structural indicator reflects the prevalence of triangle motifs, suggesting that densely inter‑connected subgraphs are more likely to correspond to real protein complexes or pathways.
A striking observation is that nearly every protein (≈ 96 % of the 5,800 proteins in the full network) belongs to at least one functionally homogeneous community at some resolution. By tracing the community membership of individual proteins across the γ spectrum, the authors illustrate how a single protein can participate in distinct functional contexts. For example, the transcription factor GAL4 is part of a broad transcription‑regulation module at low resolution but joins a small, galactose‑metabolism‑specific module at intermediate resolution. This multi‑scale perspective provides biologists with a nuanced view of protein function that is invisible when a single clustering scale is used.
The study concludes that there is no single “optimal” scale for community detection in yeast PPI networks. Instead, a range of resolution parameters yields the most biologically coherent modules, and the mean clustering coefficient can be used as a practical filter to prioritize those modules. The authors propose that this approach—systematically varying resolution, evaluating functional homogeneity with multiple independent annotations, and leveraging structural metrics—offers a robust framework for extracting biologically relevant modules from large interaction networks. Potential applications include disease‑network analysis, drug‑target identification, and comparative studies of modular evolution across species.
Comments & Academic Discussion
Loading comments...
Leave a Comment