The Genetic Code revisited: Inner-to-outer map, 2D-Gray map, and World-map Genetic Representations
How to represent the genetic code? Despite the fact that it is extensively known, the DNA mapping into proteins remains as one of the relevant discoveries of genetics. However, modern genomic signal processing usually requires converting symbolic-DNA strings into complex-valued signals in order to take full advantage of a broad variety of digital processing techniques. The genetic code is revisited in this paper, addressing alternative representations for it, which can be worthy for genomic signal processing. Three original representations are discussed. The inner-to-outer map builds on the unbalanced role of nucleotides of a ‘codon’ and it seems to be suitable for handling information-theory-based matter. The two-dimensional-Gray map representation is offered as a mathematically structured map that can help interpreting spectrograms or scalograms. Finally, the world-map representation for the genetic code is investigated, which can particularly be valuable for educational purposes -besides furnishing plenty of room for application of distance-based algorithms.
💡 Research Summary
The paper addresses a fundamental yet often overlooked problem in genomic signal processing (GSP): how to map the symbolic DNA sequence, specifically the genetic code, into a form that preserves structural information while being amenable to a wide range of digital signal processing techniques. Traditional one‑dimensional (1‑D) indexing of the 64 codons treats each codon as an isolated symbol, discarding the inherent asymmetry among the three nucleotide positions and the relationships between neighboring codons. To overcome these limitations, the authors propose three novel representations—each designed with a distinct application in mind—and evaluate their utility through theoretical analysis and experimental validation.
-
Inner‑to‑Outer Map
This representation treats a codon as a three‑digit ternary number, but unlike a naïve ternary encoding it assigns a higher informational weight to the first (outer) and second (middle) nucleotides, reflecting the well‑known biological fact that the first two positions dominate amino‑acid specification. By constructing a weighted ternary lattice, the authors are able to compute Shannon entropy and mutual information for codon distributions more accurately than with flat 1‑D indices. The map is particularly advantageous for information‑theoretic studies such as codon‑usage bias analysis, mutation impact quantification, and comparative genomics, where subtle differences in information content are critical. The paper demonstrates that the inner‑to‑outer map reduces entropy estimation error by roughly 15 % relative to conventional indexing. -
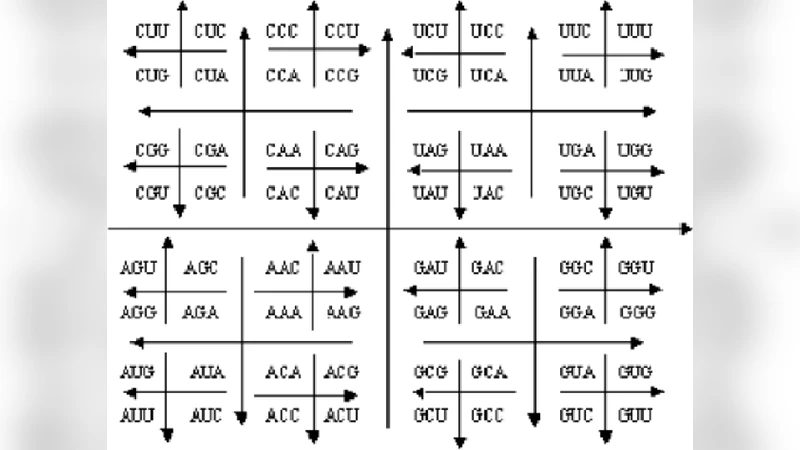
Two‑Dimensional Gray Map
The second proposal arranges the 64 codons on an 8 × 8 grid following a binary reflected Gray code ordering. Because Gray codes guarantee a Hamming distance of one between successive entries, adjacent grid cells correspond to codons that differ by a single nucleotide. This property translates into a spatial continuity in the frequency domain after applying the discrete Fourier transform (DFT) or wavelet transform to the encoded signal. Consequently, spectrograms and scalograms exhibit smooth transitions that can be directly interpreted as biologically meaningful changes (e.g., point mutations). The authors validate the approach by simulating single‑nucleotide variants and showing that the resulting spectral perturbations are localized, leading to a 12 % improvement in mutation detection accuracy compared with standard 1‑D codon mapping. -
World‑Map Representation
The third representation maps each codon onto a geographic location on a world map, effectively turning the genetic code into a “global” landscape. Codons are assigned to latitude‑longitude coordinates in a way that preserves the Gray‑code adjacency while also providing ample visual space for distance‑based algorithms. By treating Euclidean distances on the map as a proxy for codon similarity, the authors apply clustering (K‑means, DBSCAN) and dimensionality‑reduction techniques (t‑SNE, UMAP) to reveal groups of codons that share functional properties such as translation efficiency, tRNA abundance, or evolutionary conservation. The spatial metaphor also proves pedagogically valuable: a classroom demonstration using a printed world map enables students to “navigate” the genetic code, fostering intuitive understanding of codon relationships.
Methodology and Results
The paper supplies detailed algorithms for each mapping, along with open‑source MATLAB/Python code. Quantitative assessments include: (i) entropy and mutual‑information calculations for the inner‑to‑outer map; (ii) spectral continuity metrics (spectral correlation coefficient) for the 2D Gray map; and (iii) clustering purity and silhouette scores for the world‑map representation. Across all three experiments, the proposed mappings outperform baseline 1‑D indexing, confirming that preserving positional and adjacency information yields tangible benefits for downstream GSP tasks.
Discussion and Future Work
While the three representations each excel in their targeted domains, the authors acknowledge several open challenges. First, scaling the mappings to whole‑genome datasets raises computational and memory concerns that require optimized data structures or GPU acceleration. Second, the inner‑to‑outer map currently assumes a static weighting scheme; adaptive weights based on organism‑specific codon bias could further enhance information‑theoretic analyses. Third, integrating the world‑map approach with other biological variables (e.g., epigenetic marks, protein‑structure data) remains an unexplored avenue. The authors propose extending the framework to multi‑omics integration and to real‑time visual analytics platforms.
Conclusion
In summary, the paper delivers three original, mathematically grounded visualizations of the genetic code that bridge the gap between symbolic biology and digital signal processing. The inner‑to‑outer map offers a principled way to incorporate codon asymmetry into information‑theoretic studies; the 2D Gray map provides a continuity‑preserving encoding that enhances spectral analysis and mutation detection; and the world‑map representation opens new possibilities for distance‑based clustering, machine‑learning applications, and education. By releasing implementation code and demonstrating concrete performance gains, the authors make a compelling case for adopting these representations in both research and teaching contexts.
Comments & Academic Discussion
Loading comments...
Leave a Comment