High dimensional Bayesian inference for Gaussian directed acyclic graph models
In this paper, we consider Gaussian models Markov with respect to an arbitrary DAG. We first construct a family of conjugate priors for the Cholesky parametrization of the covariance matrix of such models. This family has as many shape parameters as the DAG has vertices, and naturally extends the work of Geiger and Heckerman [8]. From these distributions, we derive prior distributions for the covariance and precision parameters of the Gaussian DAG Markov models. Our works thus extends the work of Dawid and Lauritzen [5] and Letac and Massam [16] for Gaussian models Markov with respect to a decomposable graph to arbitrary DAGs. For this reason, we call our distributions DAG-Wishart distributions. An advantage of these distributions is that they possess strong hyper Markov properties and thus allow for explicit estimation of the covariance and precision parameters, regardless of the dimension of the problem. They also allow us to develop methodology for model selection and covariance estimation in the space of DAG-Markov models. We demonstrate via several numerical examples that the proposed method scales well to high-dimensions.
💡 Research Summary
This paper addresses Bayesian inference for Gaussian graphical models whose conditional independence structure is encoded by an arbitrary directed acyclic graph (DAG). While conjugate priors for Gaussian models have been extensively studied for complete graphs, decomposable undirected graphs, and for perfect DAGs (those equivalent to decomposable graphs), a general prior for arbitrary DAGs has been lacking because the space of admissible covariance or precision matrices is a curved manifold without a Lebesgue‑dominated density.
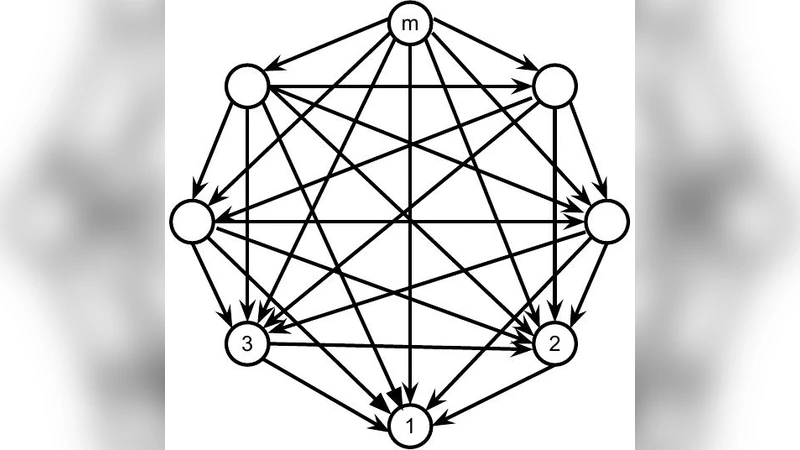
The authors propose a new family of conjugate priors, called DAG‑Wishart distributions, defined on the Cholesky parametrization of the precision matrix. For a DAG D with vertices ordered so that parents precede children, the precision matrix Ω can be uniquely written as Ω = L D⁻¹ Lᵀ, where L is a lower‑triangular matrix with unit diagonal and zeros in positions that violate the parent ordering, and D is a positive diagonal matrix. This parametrization naturally separates the model into p independent blocks, one for each vertex i, consisting of the diagonal element D_{ii} and the vector of regression coefficients L_{pa(i),i}.
Starting from the classical Wishart distribution for a complete graph, the authors map it to the Cholesky space Θ_D and compute the Jacobian of the transformation. The resulting density on Θ_D has the form
π(D,L) ∝ exp{−½ tr(L D Lᵀ U)} ∏{i=1}^p (D{ii})^{α_i/2 − |pa(i)|/2 − 1},
where U is a positive‑definite scale matrix and α_i are vertex‑specific shape parameters. In the classical Wishart case all α_i are deterministic functions of a single global shape η, but the authors deliberately treat them as independent hyper‑parameters, thereby achieving a “multi‑shape” prior.
Because the density factorizes over vertices, the DAG‑Wishart enjoys a strong hyper‑Markov property: conditional on the data, each (D_{ii}, L_{pa(i),i}) pair updates independently. Specifically, D_{ii} follows an inverse‑Gamma distribution and L_{pa(i),i} follows a multivariate normal distribution whose covariance is proportional to D_{ii}. Consequently, posterior moments can be obtained in closed form without resorting to MCMC, even when the dimension p greatly exceeds the sample size n.
To obtain priors on the original covariance (Σ) and precision (Ω) spaces, the authors use the “completion” map that uniquely reconstructs a full positive‑definite matrix from its incomplete version defined by the DAG. The Jacobian of this map is derived, allowing the DAG‑Wishart density to be transferred back to PD_D (the space of admissible covariances) and P_D (the space of admissible precisions). The resulting priors retain the multi‑shape flexibility and hyper‑Markov structure.
The paper also develops a Bayesian model‑selection framework. By placing independent Bernoulli–Beta priors on the presence of each edge (i.e., on each parent set pa(i)), the posterior probability of any DAG can be computed analytically up to a normalizing constant. The authors compare this approach with the penalized‑likelihood Lasso‑DAG method of
Comments & Academic Discussion
Loading comments...
Leave a Comment