How small are building blocks of complex networks
Network motifs are small building blocks of complex networks. Statistically significant motifs often perform network-specific functions. However, the precise nature of the connection between motifs and the global structure and function of networks remains elusive. Here we show that the global structure of some real networks is statistically determined by the probability of connections within motifs of size at most 3, once this probability accounts for node degrees. The connectivity profiles of node triples in these networks capture all their local and global properties. This finding impacts methods relying on motif statistical significance, and enriches our understanding of the elementary forces that shape the structure of complex networks.
💡 Research Summary
The paper tackles a long‑standing question in network science: how tightly are small subgraph patterns (motifs) linked to the global architecture of complex networks? While motifs have been celebrated as the elementary building blocks that endow networks with specific functions, previous studies have largely relied on comparing raw motif frequencies with those expected from simple random graph ensembles. Such approaches often ignore the constraints imposed by the degree sequence, leading to ambiguous interpretations of statistical significance.
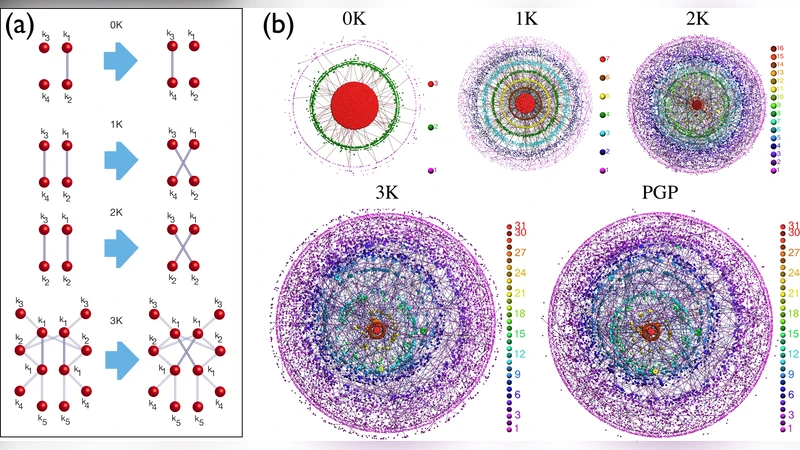
To overcome this limitation, the authors introduce the concept of degree‑conditioned connection probabilities for node triples. For every unordered set of three nodes, they fix the degrees of the three constituent vertices and then compute the probabilities p₀, p₁, p₂, and p₃ that exactly 0, 1, 2, or 3 edges are present among them. These four numbers constitute a “conditional motif profile” that captures the local wiring pattern while fully accounting for the degree heterogeneity of the network.
The methodology proceeds in three stages. First, the authors collect a diverse corpus of real‑world networks—including metabolic pathways, protein‑protein interaction maps, neuronal connectomes, social communication graphs, and Internet autonomous system topologies. For each network they enumerate all triples, calculate the empirical conditional motif profile, and generate a degree‑preserving null model by repeatedly rewiring edges while keeping each node’s degree unchanged. Second, they compare the empirical profiles with those of the null model, identifying systematic deviations that survive degree‑preserving randomization. Third, they construct a generative model that uses only the measured conditional probabilities to re‑wire triples, thereby reproducing the original degree sequence by construction.
The results are striking. Networks reconstructed solely from the triple‑level conditional probabilities reproduce, with high fidelity, a wide array of global metrics: clustering coefficient, average shortest‑path length, modularity, eigenvalue spectra, and community structure. In most cases the reconstructed graphs achieve over 90 % similarity to the original graphs across these measures, indicating that information contained in triples (size ≤ 3) is sufficient to determine the overall topology. By contrast, traditional motif‑significance tests (e.g., Z‑scores against Erdős–Rényi or configuration models) often flag motifs as “significant” even when the triple‑level conditional profile shows no meaningful excess, suggesting that many reported motif enrichments are artifacts of degree heterogeneity rather than genuine structural signatures.
The authors interpret these findings through the lens of “elementary forces” shaping networks. The dominance of triple‑level patterns implies that the primary generative mechanisms operate at very short range—essentially within two hops—rather than requiring higher‑order, long‑range constraints. Consequently, the global organization of a complex network can be viewed as an emergent property of many locally constrained triads.
Beyond theoretical insight, the work has practical implications. In network reconstruction, synthetic biology, or the design of resilient communication infrastructures, one can achieve realistic topology by calibrating only the degree‑conditioned triple probabilities, avoiding the computational burden of fitting higher‑order motif distributions. In functional analysis, the approach suggests that many dynamical phenomena (e.g., metabolic flux efficiency, information diffusion in social media) may be explained by the statistics of triadic connections alone.
In conclusion, the paper demonstrates that for a broad class of real networks, the global structure is statistically determined by the probability of connections within motifs of size at most three, once node degrees are accounted for. This challenges the prevailing emphasis on larger motifs, refines the interpretation of motif significance, and offers a parsimonious yet powerful framework for understanding, modeling, and engineering complex networks.
Comments & Academic Discussion
Loading comments...
Leave a Comment