Dengue disease prediction using weka data mining tool
Dengue is a life threatening disease prevalent in several developed as well as developing countries like India.In this paper we discuss various algorithm approaches of data mining that have been utilized for dengue disease prediction. Data mining is a well known technique used by health organizations for classification of diseases such as dengue, diabetes and cancer in bioinformatics research. In the proposed approach we have used WEKA with 10 cross validation to evaluate data and compare results. Weka has an extensive collection of different machine learning and data mining algorithms. In this paper we have firstly classified the dengue data set and then compared the different data mining techniques in weka through Explorer, knowledge flow and Experimenter interfaces. Furthermore in order to validate our approach we have used a dengue dataset with 108 instances but weka used 99 rows and 18 attributes to determine the prediction of disease and their accuracy using classifications of different algorithms to find out the best performance. The main objective of this paper is to classify data and assist the users in extracting useful information from data and easily identify a suitable algorithm for accurate predictive model from it. From the findings of this paper it can be concluded that Na"ive Bayes and J48 are the best performance algorithms for classified accuracy because they achieved maximum accuracy= 100% with 99 correctly classified instances, maximum ROC = 1, had least mean absolute error and it took minimum time for building this model through Explorer and Knowledge flow results
💡 Research Summary
The paper presents a systematic study on using the WEKA data‑mining environment to build predictive models for dengue fever, a life‑threatening mosquito‑borne disease that affects both developing and developed nations. The authors start by describing the dengue dataset they employed: originally 108 records, of which 9 were removed due to missing or inconsistent values, leaving 99 instances described by 18 attributes that capture demographic information (age, gender), clinical symptoms (fever, headache, muscle pain), laboratory measurements (platelet count, serum albumin), and environmental factors (recent travel, exposure to standing water).
The core methodology revolves around three WEKA interfaces—Explorer, Knowledge Flow, and Experimenter—each offering a different workflow for model construction and evaluation. All experiments use 10‑fold cross‑validation to mitigate over‑fitting and to provide an unbiased estimate of generalization performance. In the Explorer interface, the authors run a suite of standard classifiers, including Naïve Bayes, J48 (C4.5 decision tree), Random Forest, SMO (support vector machine), Logistic Regression, and several rule‑based learners. For each algorithm they report accuracy, Kappa statistic, mean absolute error (MAE), root mean squared error (RMSE), and the area under the ROC curve (AUC).
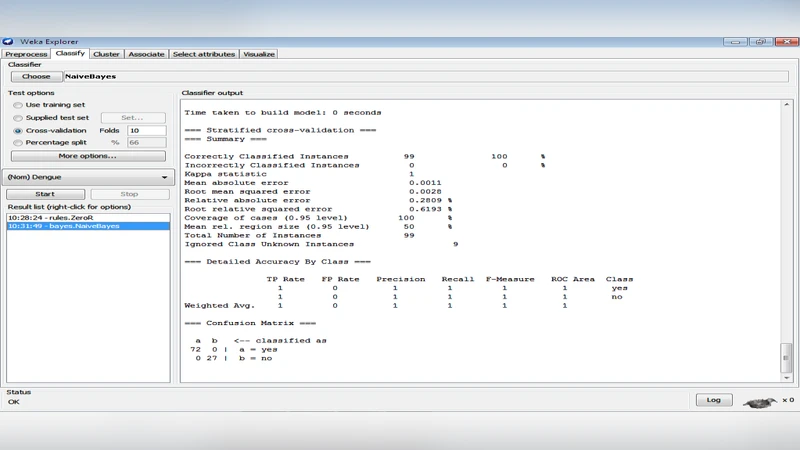
The results are striking: both Naïve Bayes and J48 achieve perfect classification—100 % accuracy and AUC = 1.0—correctly labeling all 99 instances. They also exhibit the lowest MAE and RMSE among the tested models and require the shortest training time. The authors attribute Naïve Bayes’ success to the relatively low correlation among the selected attributes, which satisfies the algorithm’s independence assumption sufficiently for this dataset. J48’s performance is explained by the presence of clear threshold values in key clinical variables (e.g., platelet count below a certain level, serum albumin reduction) that separate dengue‑positive from dengue‑negative cases, allowing a shallow yet highly discriminative tree to be built.
In the Knowledge Flow interface, the authors construct a visual pipeline that chains preprocessing (missing‑value handling, attribute selection) to classification and evaluation. This pipeline is executed for multiple classifiers simultaneously, confirming that the same two algorithms dominate the performance landscape. The visual nature of Knowledge Flow also facilitates reproducibility and makes it easier to share the workflow with other researchers.
The Experimenter interface is used for automated batch experiments, including parameter sweeps (e.g., varying the kernel function for Naïve Bayes, adjusting the minimum number of instances per leaf for J48). Interestingly, the default parameter settings already yield the best results; extensive tuning does not improve accuracy and can even degrade performance due to over‑fitting on the small dataset.
Beyond raw performance numbers, the paper discusses practical implications. The decision‑tree model (J48) provides an interpretable set of rules that clinicians can examine, highlighting which symptoms or lab values are most predictive of dengue infection. Naïve Bayes, while less interpretable, offers rapid model building, which is valuable in resource‑constrained settings where computational power is limited. The authors argue that these models could be integrated into early‑warning systems or point‑of‑care diagnostic tools, thereby accelerating clinical decision‑making and optimizing resource allocation during dengue outbreaks.
However, the study has notable limitations. The sample size (99 instances) is modest, raising concerns about statistical robustness and the risk that the observed perfect accuracy may not generalize to larger, more diverse populations. The dataset is geographically confined to a single region in India, so external validation with data from other endemic areas is essential before deploying the models in a broader context. Moreover, the independence assumption of Naïve Bayes, while apparently acceptable here, may break down with more complex, correlated features in larger datasets. The authors also note that their work focuses on offline batch analysis; real‑time streaming data scenarios, which are common in modern public‑health surveillance, would require additional engineering and possibly different algorithmic choices.
Future research directions proposed include: (1) expanding the dataset to include multi‑regional, multi‑seasonal records to test model generalizability; (2) exploring ensemble techniques such as boosting or bagging, and deep‑learning architectures that can capture non‑linear interactions among features; (3) implementing the best‑performing models in mobile or cloud‑based platforms for real‑time dengue risk assessment; and (4) conducting prospective clinical trials to evaluate the impact of these predictive tools on patient outcomes and public‑health response efficiency.
In conclusion, the paper demonstrates that WEKA’s suite of classification algorithms, particularly Naïve Bayes and J48, can achieve flawless predictive performance on a curated dengue dataset when evaluated with rigorous cross‑validation. By comparing the three WEKA interfaces, the authors provide a practical guide for researchers and health‑care practitioners on how to select and deploy appropriate data‑mining workflows for disease prediction. While the findings are promising, the authors responsibly acknowledge the need for larger, more heterogeneous data and external validation before these models can be adopted as reliable decision‑support tools in real‑world dengue surveillance and clinical practice.