Transcriptomic and metabolomic analysis of copper stress acclimation in Ectocarpus siliculosus highlights signaling and tolerance mechanisms in brown algae
Brown algae are sessile macro-organisms of great ecological relevance in coastal ecosystems. They evolved independently from land plants and other multicellular lineages, and therefore hold several original ontogenic and metabolic features. Most brown algae grow along the coastal zone where they face frequent environmental changes, including exposure to toxic levels of heavy metals such as copper (Cu). We carried out large-scale transcriptomic and metabolomic analyses to decipher the short-term acclimation of the brown algal model E. siliculosus to Cu stress, and compared these data to results known for other abiotic stressors. This comparison demonstrates that Cu induces oxidative stress in E. siliculosus as illustrated by the transcriptomic overlap between Cu and H2O2 treatments. The common response to Cu and H2O2 consisted in the activation of the oxylipin and the repression of inositol signaling pathways, together with the regulation of genes coding for several transcription-associated proteins. Concomitantly, Cu stress specifically activated a set of genes coding for orthologs of ABC transporters, a P1B-type ATPase, ROS detoxification systems such as a vanadium-dependent bromoperoxidase, and induced an increase of free fatty acid contents. Finally we observed, as a common abiotic stress mechanism, the activation of autophagic processes on one hand and the repression of genes involved in nitrogen assimilation on the other hand. Comparisons with data from green plants indicate that some processes involved in Cu and oxidative stress response are conserved across these two distant lineages. At the same time the high number of yet uncharacterized brown alga-specific genes induced in response to copper stress underlines the potential to discover new components and molecular interactions unique to these organisms. Of particular interest for future research is the potential cross-talk between reactive oxygen species (ROS)-, myo-inositol-, and oxylipin signaling.
💡 Research Summary
This study investigates the short‑term acclimation of the brown alga Ectocarpus siliculosus to copper (Cu) stress by integrating large‑scale transcriptomic and metabolomic analyses and comparing the results with those obtained for other abiotic stressors. Algae were exposed to 10 µM Cu for 24 h, after which RNA‑Seq was performed to identify differentially expressed genes (DEGs) (false‑discovery‑rate < 0.05, |log2FC| > 1). Metabolite profiling was carried out using GC‑MS and LC‑MS, quantifying over 150 compounds including free fatty acids, sterols, amino acids, and sugars.
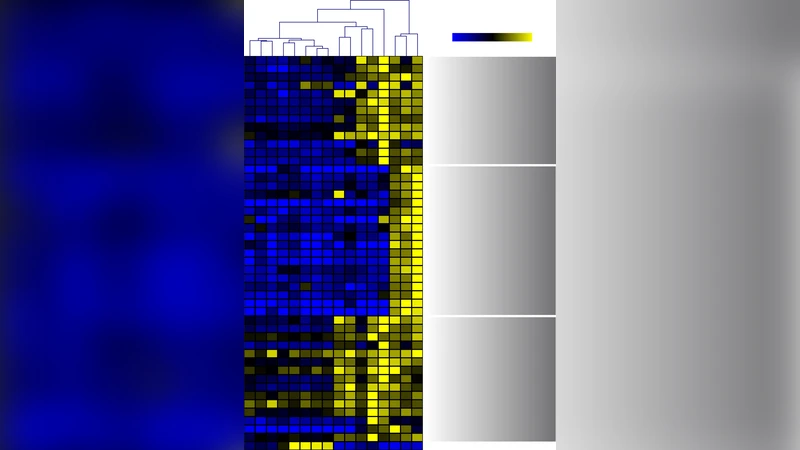
A striking finding is the extensive overlap between the Cu‑responsive transcriptome and that induced by hydrogen peroxide (H₂O₂), indicating that Cu toxicity is largely mediated by oxidative stress. Genes involved in oxylipin biosynthesis and signaling were up‑regulated in both treatments, with notable increases in 13‑hydroxy‑octadecatrienoic acid (13‑HOTrE) and 12‑hydroxy‑eicosatetraenoic acid (12‑HETE). Conversely, components of the myo‑inositol signaling pathway were down‑regulated, suggesting a shift in calcium and ROS homeostasis.
Cu‑specific responses included strong induction of several ATP‑binding cassette (ABC) transporters, particularly members of the ABCC and ABCG subfamilies, and a P1B‑type ATPase known to export Cu⁺ ions. A vanadium‑dependent bromoperoxidase (VBPO) was also highly expressed; this enzyme is characteristic of brown algae and contributes to halogenation reactions and reactive oxygen species (ROS) detoxification. Metabolically, free fatty acids such as palmitic and oleic acids accumulated markedly, providing substrates for membrane remodeling and oxylipin production.
Across all abiotic stresses examined, two conserved mechanisms emerged: activation of autophagy (up‑regulation of ATG8, ATG12, and related genes) and repression of nitrogen assimilation genes (glutamine synthetase, asparagine synthetase). These responses likely reflect a cellular strategy to conserve energy and nitrogen under adverse conditions.
Comparative analysis with terrestrial plants revealed that while some ROS‑related detoxification genes are conserved, a large proportion of the Cu‑responsive genes are brown‑alga‑specific and remain functionally uncharacterized. This underscores the evolutionary divergence of brown algae and points to a rich source of novel stress‑response components. The authors highlight the potential cross‑talk among ROS, myo‑inositol, and oxylipin signaling pathways as a promising area for future research.
In summary, copper exposure triggers oxidative stress in E. siliculosus, leading to a coordinated transcriptional reprogramming that includes metal export (ABC transporters, P1B‑ATPase), specialized ROS‑scavenging enzymes (VBPO), lipid‑derived signaling (oxylipins), autophagic turnover, and down‑regulation of nitrogen metabolism. While some aspects of this response are shared with green plants, many elements are unique to brown algae, reflecting their distinct evolutionary history and ecological niche. The study provides a comprehensive resource for understanding metal tolerance in marine macroalgae and lays the groundwork for dissecting the molecular mechanisms underlying ROS‑myo‑inositol‑oxylipin cross‑communication.
Comments & Academic Discussion
Loading comments...
Leave a Comment