Aggregation-fragmentation model of robust concentration gradient formation
Concentration gradients of signaling molecules are essential for patterning during development and they have been observed in both unicellular and multicellular systems. In subcellular systems, clustering of the signaling molecule has been observed. We develop a theoretical model of cluster- mediated concentration gradient formation based on the Becker-Doring equations of aggregation- fragmentation processes. We show that such a mechanism produces robust concentration gradients on realistic time and spatial scales so long as the process of clustering does not significantly stabilize the signaling molecule. Finally, we demonstrate that such a model is applicable to the pom1p subcellular gradient in fission yeast.
💡 Research Summary
In this paper the authors develop a quantitative theory for how subcellular concentration gradients can be generated and maintained when the signaling molecules form transient clusters on the plasma membrane. Starting from the classic Becker‑Döring framework for aggregation‑fragmentation kinetics, they write down a set of reaction‑diffusion equations for monomers (n₁) and clusters of size s (n_s). The key mechanistic assumptions, motivated by live‑cell imaging of the fission‑yeast protein Pom1p, are: (i) clusters grow and shrink by the addition or loss of single monomers only; (ii) clusters do not detach from the membrane as whole entities; (iii) new molecules are inserted only as monomers at the source.
Diffusion coefficients are taken to scale as D_s = D₁ s⁻ʳᴰ (with r_D≈1), reflecting experimental observations that larger clusters move more slowly. The membrane‑dissociation rate is μ_s = μ₁ s⁻ʳµ, where r_µ=0 corresponds to no stabilization of the molecule inside a cluster, while r_µ=1 would mean that clustering dramatically prolongs the molecule’s lifetime. The authors show analytically that only the former case yields robust gradients.
Through dimensional analysis the originally large parameter space collapses to seven dimensionless numbers. Two of them, κ = J α₁/(√D₁ μ₁³) and β̃ = β₁/μ₁, control respectively the balance between insertion/aggregation versus diffusion/dissociation, and the relative speed of fragmentation. Using experimentally measured values for Pom1p they obtain κ≈425 and β̃≈20, a regime where clusters form but do not dominate the dynamics far from the source.
Numerical integration (MATLAB pde45 and bvp4c) yields steady‑state total concentration profiles N_T(x)=∑_s s n_s(x) that decay sharply near the source and flatten at larger distances. Large clusters are confined to the source region, while smaller clusters populate the distal zone. An effective diffusion coefficient D_eff(x)=∑_s s n_s D_s / N_T(x) rises with distance, indicating that clustering locally suppresses diffusion near the source and enhances it farther away—a built‑in diffusion‑modulation mechanism.
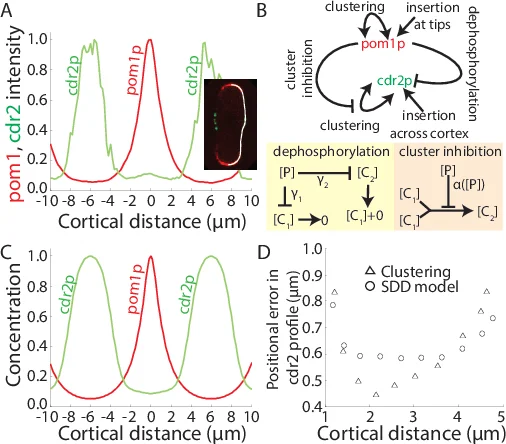
The authors then assess robustness. First, they examine how fluctuations in the insertion rate J translate into positional error δx = δρ/|∂ρ/∂x| at a given concentration threshold. Compared with a simple linear diffusion‑degradation (SDD) model and a non‑linear degradation (NLD) model, the clustering model (with r_µ=0) shows comparable or smaller δx, i.e. it is equally robust to source‑strength variations. Second, they consider intrinsic stochasticity modeled as Poisson noise (δρ_int ∝ √ρ). Because D_eff increases with distance, time‑averaging is more effective in the distal region, reducing the impact of intrinsic fluctuations. The clustering model is slightly less robust than the CDD (concentration‑dependent diffusion) model but still outperforms the NLD model.
Finally, the theory is applied to the Pom1p gradient in fission yeast. Using the measured cluster size distribution and kinetic rates, the model reproduces the experimentally observed concentration profile and the spatial segregation of large versus small clusters. This quantitative agreement validates the hypothesis that Pom1p’s subcellular gradient is shaped by a balance of monomer insertion, cluster‑mediated aggregation/fragmentation, and size‑dependent diffusion, without requiring strong stabilization of the protein within clusters.
Overall, the study demonstrates that transient clustering can act as a self‑tuning diffusion regulator, generating concentration gradients that are both fast to establish (seconds) and robust to biologically relevant fluctuations. The work provides a mechanistic bridge between observed molecular clustering and morphogen‑like gradient formation, and suggests that engineering similar aggregation‑fragmentation modules could be a powerful strategy for synthetic biology applications that demand precise spatial patterning.
Comments & Academic Discussion
Loading comments...
Leave a Comment