Can bio-inspired information processing steps be realized as synthetic biochemical processes?
We consider possible designs and experimental realiza-tions in synthesized rather than naturally occurring bio-chemical systems of a selection of basic bio-inspired information processing steps. These include feed-forward loops, which have been identified as the most common information processing motifs in many natural pathways in cellular functioning, and memory-involving processes, specifically, associative memory. Such systems should not be designed to literally mimic nature. Rather, we can be guided by nature’s mechanisms for experimenting with new information/signal processing steps which are based on coupled biochemical reactions, but are vastly simpler than natural processes, and which will provide tools for the long-term goal of understanding and harnessing nature’s information processing paradigm. Our biochemical processes of choice are enzymatic cascades because of their compatibility with physiological processes in vivo and with electronics (e.g., electrodes) in vitro allowing for networking and interfacing of enzyme-catalyzed processes with other chemical and biochemical reactions. In addition to designing and realizing feed-forward loops and other processes, one has to develop approaches to probe their response to external control of the time-dependence of the input(s), by measuring the resulting time-dependence of the output. The goal will be to demonstrate the expected features, for example, the delayed response and stabilizing effect of the feed-forward loops.
💡 Research Summary
The paper explores the feasibility of reproducing fundamental bio‑inspired information‑processing steps in wholly synthetic biochemical systems, focusing on enzymatic cascades as the implementation platform. The authors deliberately avoid attempting a literal recreation of natural pathways; instead, they extract the essential functional motifs—feed‑forward loops (FFLs) and associative memory—and redesign them as minimal, controllable enzyme‑based circuits that can be interfaced with electronic measurement tools.
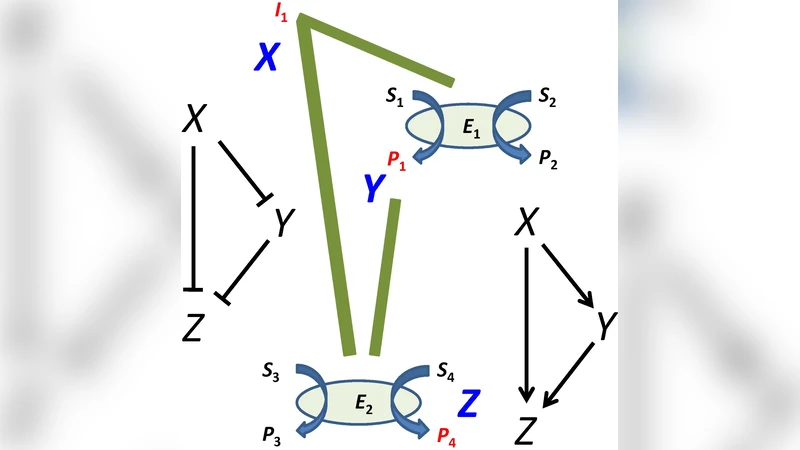
The first major contribution is the design and experimental realization of a coherent type‑1 feed‑forward loop using three enzymes. Substrate S is converted by enzyme E1 into an intermediate X, which activates enzyme E2 to produce the final output Z. In parallel, S is also directly converted to Z by enzyme E3, establishing the classic dual‑path architecture of an FFL. By tuning kinetic parameters (Km, Vmax), reaction conditions (pH, temperature), and enzyme immobilization density on an electrode surface, the authors achieve a controlled delay between input changes and output response, as well as a pronounced attenuation of stochastic input fluctuations. The loop’s stabilizing effect is quantified by measuring the reduction in output variance under noisy input conditions.
The second contribution is a synthetic associative‑memory module. Two distinct inputs, A and B, must be present simultaneously to trigger the formation of a heterodimeric enzyme complex C·D. This complex initiates a self‑reinforcing feedback loop that continues to generate output Z even after A and B are removed, thereby embodying a short‑term memory trace. The duration of memory retention is governed by the dissociation constant of the complex and the catalytic turnover of the feedback enzymes; by adjusting these parameters, the system can be switched between “brief” (≈5 min) and “extended” (up to 20 min) memory regimes.
Experimentally, the circuits are embedded in a microfluidic platform that delivers precisely programmed concentration waveforms of the inputs. Enzymes are covalently attached to gold electrodes, allowing the authors to monitor reaction progress in real time via amperometric detection. The current‑time traces provide a direct readout of the system’s dynamic behavior: the onset of delayed output in the FFL, the magnitude of noise suppression, and the persistence of output after the associative cue is withdrawn. Computational modeling using ordinary differential equations corroborates the experimental data, confirming that the designed kinetic couplings reproduce the expected theoretical behavior.
Key insights emerging from the study include: (1) enzymatic cascades naturally embody non‑linear dynamics, diffusion constraints, and environmental sensitivities, making them well‑suited for implementing complex temporal processing without the need for external power beyond the chemical substrates; (2) the combination of feed‑forward control and feedback‑based memory creates a primitive “chemical neural network” capable of filtering transient signals while retaining salient patterns; (3) the electrode‑based detection scheme offers a seamless bridge between biochemical computation and electronic readout, opening pathways toward hybrid bio‑electronic devices, smart drug‑delivery platforms, and synthetic cells that can process information in a manner reminiscent of living organisms.
In conclusion, the authors demonstrate that (i) a synthetic feed‑forward loop can be constructed from a minimal set of enzymes and exhibits the hallmark delayed response and noise‑filtering properties of its natural counterpart, and (ii) an associative‑memory circuit can be realized that maintains output after the conditioning inputs disappear. These proof‑of‑concept systems lay the groundwork for scaling up to larger, more intricate enzyme networks, ultimately advancing the field of bio‑inspired computing toward practical, programmable biochemical processors.
Comments & Academic Discussion
Loading comments...
Leave a Comment