ComPPI, a cellular compartment-specific database for protein-protein interaction network analysis
Here we present ComPPI, a cellular compartment specific database of proteins and their interactions enabling an extensive, compartmentalized protein-protein interaction network analysis (http://ComPPI.LinkGroup.hu). ComPPI enables the user to filter biologically unlikely interactions, where the two interacting proteins have no common subcellular localizations and to predict novel properties, such as compartment-specific biological functions. ComPPI is an integrated database covering four species (S. cerevisiae, C. elegans, D. melanogaster and H. sapiens). The compilation of nine protein-protein interaction and eight subcellular localization data sets had four curation steps including a manually built, comprehensive hierarchical structure of more than 1600 subcellular localizations. ComPPI provides confidence scores for protein subcellular localizations and protein-protein interactions. ComPPI has user-friendly search options for individual proteins giving their subcellular localization, their interactions and the likelihood of their interactions considering the subcellular localization of their interacting partners. Download options of search results, whole proteomes, organelle-specific interactomes and subcellular localization data are available on its website. Due to its novel features, ComPPI is useful for the analysis of experimental results in biochemistry and molecular biology, as well as for proteome-wide studies in bioinformatics and network science helping cellular biology, medicine and drug design.
💡 Research Summary
The paper introduces ComPPI, a web‑based database that integrates protein‑protein interaction (PPI) data with subcellular localization information to enable compartment‑specific network analysis. The authors compiled nine major PPI resources (BioGRID, IntAct, DIP, MINT, HPRD, MIPS, BIND, HINT, STRING) and eight localization datasets (UniProt, Gene Ontology, Human Protein Atlas, Compartments, LOCATE, SubCellBarCode, Cell Atlas, PSORTdb) for four model organisms: Saccharomyces cerevisiae, Caenorhabditis elegans, Drosophila melanogaster, and Homo sapiens. After extensive curation—including duplicate removal, manual conflict resolution, and source‑specific weighting—the data were merged using UniProtKB accessions as a common identifier.
A distinctive feature of ComPPI is its hierarchical ontology of more than 1 600 subcellular compartments, organized in a three‑level tree (organelle → sub‑organelle → fine‑grained location). This structure allows the authors to capture both broad and highly specific localization contexts, which are essential for accurate confidence scoring.
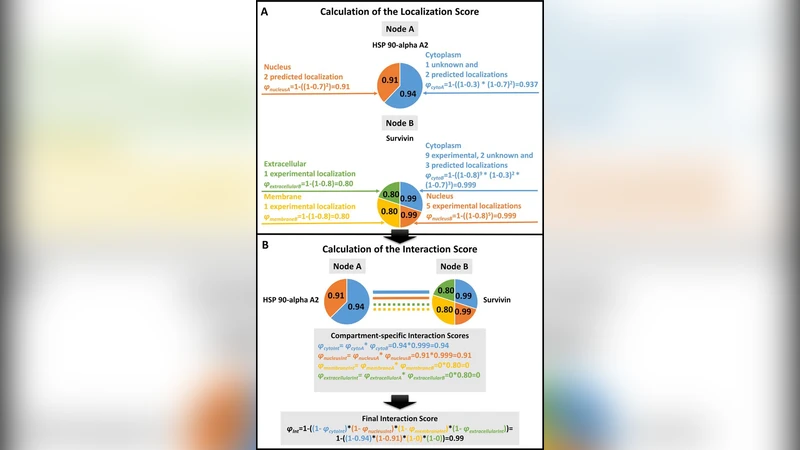
Two quantitative scores are provided for each protein and each interaction. The Localization Score (LScore) aggregates evidence from all localization sources, weighting each source by its curated reliability and by the frequency of reporting, yielding a normalized value between 0 and 1. The Interaction Score (IScore) evaluates the likelihood that two proteins can physically interact in vivo: it first identifies the set of shared compartments, multiplies the corresponding LScores, and then incorporates additional factors such as the experimental method (e.g., Y2H, co‑IP, AP‑MS) and reproducibility across studies. The resulting IScore also ranges from 0 to 1 and can be used as a filter to discard biologically implausible interactions (i.e., those lacking any common compartment).
The web interface offers a simple search box, detailed protein pages, and interactive network visualizations. Users can retrieve a protein’s full localization profile, its list of interaction partners, and the IScore for each partner. Advanced filters enable extraction of compartment‑specific interactomes (e.g., only mitochondrial or Golgi networks) and allow users to set custom IScore thresholds. All results can be downloaded in CSV or JSON format, and bulk downloads of whole‑organism proteomes, organelle‑specific interactomes, and the complete localization matrix are also available.
The authors demonstrate the utility of ComPPI through several case studies. First, they show that filtering by IScore removes a substantial fraction of false‑positive interactions that would otherwise be retained in conventional PPI datasets. Second, they illustrate how compartment‑restricted subnetworks can reveal functional modules, such as nuclear transcription complexes, that are invisible in the global network. Third, they apply the resource to disease‑related proteins, uncovering a pronounced enrichment of certain pathologies in specific organelles, thereby suggesting novel drug‑targeting strategies.
Limitations acknowledged include the restriction to four species and the dependence on experimentally verified localization data, which leaves many proteins with low LScores due to sparse evidence. The authors propose future extensions that will incorporate additional model organisms, integrate high‑resolution spatial proteomics techniques (e.g., LOPIT‑MS, hyper‑LOPIT), and employ machine‑learning models to predict missing localizations, thereby improving both coverage and scoring accuracy.
In summary, ComPPI provides a rigorously curated, compartment‑aware PPI resource that enhances the biological relevance of interaction networks, supports hypothesis generation for functional genomics, and offers a valuable platform for biomedical research, including drug discovery and systems‑level disease modeling.
Comments & Academic Discussion
Loading comments...
Leave a Comment