A consistent model for cardiac deformation estimation under abnormal ventricular muscle conditions
Deformation modeling of cardiac muscle is an important issue in the field of cardiac analysis. For this reason, many approaches have been developed to best estimate the cardiac muscle deformation, and to obtain a practical model to use in diagnostic procedures. But there are some conditions, like in case of myocardial infarction, in which the regular modeling approaches are not useful. In this section, using a point-wise approach in deformation estimation, we try to estimate the deformation under some abnormal conditions of cardiac muscle. First, the endocardial and epicardial contour points are ordered with respect to the center of gravity of endocardial contour and boundary point displacement vectors are extracted. Then to solve the governing equation of deformation, which is an elliptic equation, we apply boundary conditions in accordance with the computed displacement vectors and then the Finite Element method (FEM) will be used to solve the governing equation. Using obtained displacement field through the cardiac muscle, strain map is extracted to show the mechanical behavior of cardiac muscle. To validate the proposed algorithm in case of infracted muscle, a non-homogeneous ring is modeled using ANSYS under a uniform time varying internal pressure, which is the case in real cardiac muscle deformation and then the proposed algorithm implemented in MATLAB and the results for such problem are extracted.
💡 Research Summary
The paper addresses a critical gap in cardiac biomechanics: the inability of conventional deformation models to accurately capture the mechanical behavior of myocardium under pathological conditions such as myocardial infarction, where tissue properties become highly heterogeneous. To overcome this, the authors propose a point‑wise deformation estimation framework that begins with the extraction of endocardial and epicardial contours from cardiac images. By computing the centroid of the endocardial contour and ordering all contour points angularly around this centroid, a consistent one‑dimensional parametrization of the myocardial wall is obtained. This parametrization enables the precise calculation of displacement vectors for each point as the heart experiences a time‑varying internal pressure that mimics the physiological systolic‑diastolic cycle.
These displacement vectors are then imposed as boundary conditions on the governing elliptic partial differential equation that describes linear elastic deformation. The equation is solved using the Finite Element Method (FEM) implemented in MATLAB. Crucially, the mesh is constructed to reflect spatially varying material properties: regions representing infarcted tissue are assigned a markedly lower Young’s modulus than healthy myocardium, thereby embedding the non‑homogeneous stiffness distribution directly into the numerical model. Solving the FEM system yields a full displacement field across the myocardial wall, from which strain tensors are derived. The resulting strain map visualizes the mechanical contrast between normal and damaged zones, offering a potential diagnostic indicator for clinicians.
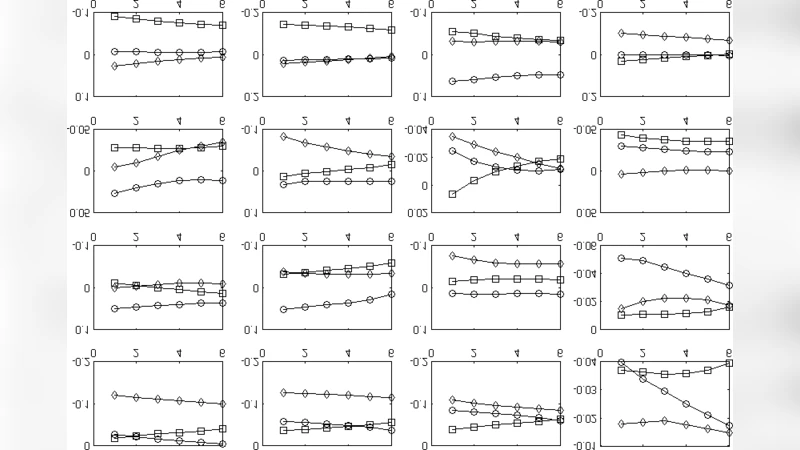
For validation, the authors build a non‑homogeneous ring model in ANSYS. The ring is subjected to a uniform, linearly increasing internal pressure, reproducing the loading conditions of a beating ventricle. A segment of the ring is designated as infarcted by reducing its elastic modulus, creating a realistic heterogeneity. The same pressure history is fed into the MATLAB algorithm, which processes the synthetic contour data, applies the point‑wise boundary conditions, and solves the FEM problem. Comparative analysis shows a high degree of agreement between the ANSYS reference solution and the proposed MATLAB results in terms of both displacement amplitudes and spatial strain distribution. This concordance demonstrates that the method can accurately recover deformation fields even when material properties vary sharply within the domain.
Key strengths of the work include: (1) the innovative use of angular ordering around the endocardial centroid to achieve a robust point‑wise correspondence between frames; (2) the direct translation of measured displacements into physically meaningful boundary conditions, preserving the causality of pressure‑driven deformation; (3) the integration of heterogeneous material modeling within a standard FEM pipeline, enabling realistic simulation of infarcted myocardium; and (4) thorough validation against a high‑fidelity commercial finite‑element solver.
Nevertheless, the study has limitations. The current implementation operates on a 2‑D planar representation of the ventricular wall, which cannot fully capture the three‑dimensional curvature and torsional dynamics of the real heart. Moreover, accurate assignment of regional material properties relies on prior knowledge or invasive measurements, potentially limiting clinical translation. Future research directions suggested by the authors include extending the point‑wise ordering to full 3‑D surface meshes derived from MRI or CT, coupling the framework with non‑invasive elastography techniques to estimate regional stiffness in vivo, and incorporating non‑linear constitutive models to better reflect the myocardium’s complex, time‑dependent behavior. By addressing these extensions, the proposed methodology could evolve into a powerful tool for patient‑specific cardiac mechanics assessment and aid in the diagnosis, monitoring, and treatment planning of myocardial diseases.