Assembling biological boolean networks using manually curated databases and prediction algorithms
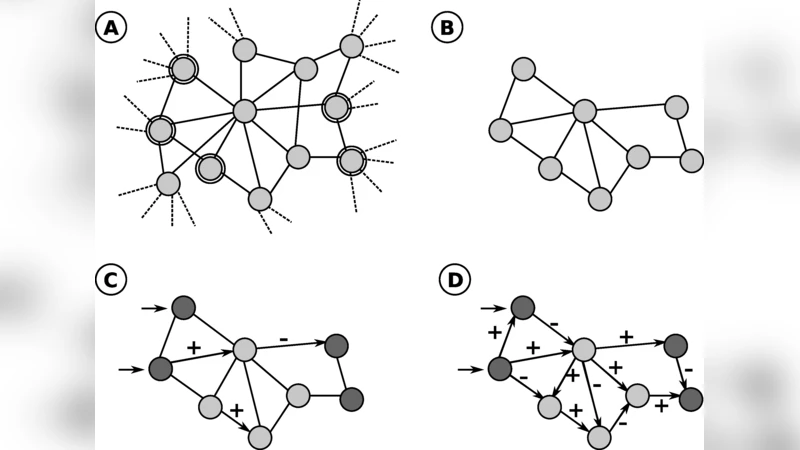
Despite the large quantity of information available, thorough researches in various biological databases are still needed in order to reconstruct and understand the steps that lead to known or new phenomena. By using protein-protein interaction networks and algorithms to extract relevant interconnections among proteins of interest, it is possible to assemble subnetworks from global interactomes. Using these extracted networks it is possible to use algorithms to predict signal directions while activation and inhibition effects can be predicted using RNA interference screenings. The result of this approach is the automatic generation of boolean networks. This way of modelling dynamical systems allows the discovery of steady states and the prediction of stimuli response.
💡 Research Summary
The paper presents an integrated workflow for constructing Boolean models of biological signaling networks by combining manually curated interaction databases with computational prediction algorithms. First, the authors aggregate protein‑protein interaction (PPI) data from major public resources such as STRING, BioGRID, and IntAct, and then apply a rigorous manual curation step that incorporates experimental validation, literature evidence, and confidence weighting to produce a high‑quality interaction set. From this global interactome, a subnetwork relevant to a user‑defined protein list (e.g., disease‑associated proteins or pathway components) is extracted using graph‑theoretic methods—minimum spanning trees, community detection, and a custom relevance score that captures both physical binding and functional association.
To endow the subnetwork with directionality, the authors train a Bayesian‑fuzzy inference model on known signaling routes (MAPK, PI3K‑AKT, NF‑κB, etc.). The model assigns probabilistic edge directions and provides confidence intervals for each prediction. Activation or inhibition signs are inferred from large‑scale RNAi (or CRISPR) screening data: the effect of gene knock‑down on downstream protein activity is quantified, statistically validated, and encoded as “+” (activation) or “‑” (inhibition) on each directed edge.
With a fully annotated directed graph, the workflow automatically translates the network into Boolean logic functions. Each node is binary (ON/OFF) and its update rule is constructed from the logical combination (AND, OR, NOT) of its upstream regulators, respecting the signed edges. The resulting Boolean model can be imported into existing simulators such as GINsim, BoolNet, or CellNetAnalyzer. The authors demonstrate that the model reproduces known steady‑states and dynamic responses to stimuli such as epidermal growth factor (EGF) and tumor necrosis factor‑α (TNF‑α), correctly predicting cell‑cycle progression, apoptosis induction, and inflammatory signaling. Moreover, the model generates novel hypotheses—for example, an indirect inhibitory circuit linking a previously uncharacterized phosphatase to the MAPK cascade—that can be experimentally tested.
The pipeline is modular: (1) data collection and manual curation, (2) subnetwork extraction, (3) direction and sign prediction, (4) Boolean model generation, and (5) simulation and analysis. Each module can be swapped or customized, and the authors release the entire framework as open‑source software, encouraging community extension and validation. In summary, the study delivers a reproducible, automated method for converting curated interactome data into Boolean dynamical models, facilitating the discovery of steady‑states, stimulus‑response behavior, and new regulatory motifs in complex biological systems.
Comments & Academic Discussion
Loading comments...
Leave a Comment