A Parallel Genetic Algorithm for Three Dimensional Bin Packing with Heterogeneous Bins
This paper presents a parallel genetic algorithm for three dimensional bin packing with heterogeneous bins using Hadoop Map-Reduce framework. The most common three dimensional bin packing problem which packs given set of boxes into minimum number of equal sized bins is proven to be NP Hard. The variation of three dimensional bin packing problem that allows heterogeneous bin sizes and rotation of boxes is computationally more harder than common three dimensional bin packing problem. The proposed Map-Reduce implementation helps to run the genetic algorithm for three dimensional bin packing with heterogeneous bins on multiple machines parallely and computes the solution in relatively short time.
💡 Research Summary
The paper introduces a parallel genetic algorithm (GA) designed to solve the three‑dimensional bin packing problem with heterogeneous bins (3D‑BPP‑H) and allows box rotations. After establishing that the classic 3D bin packing problem with identical bins is NP‑hard, the authors argue that introducing bins of varying dimensions and permitting six possible orientations for each box dramatically enlarges the search space, making the problem even more computationally demanding. To address this, they adopt a GA because of its global search capability and flexibility in handling complex constraints.
Each individual (chromosome) encodes three pieces of information: the order in which boxes are placed, a rotation code (0‑5) for each box, and the index of the bin to which the box is assigned. The fitness function combines three objectives: (1) minimizing the total number of bins used (primary goal), (2) maximizing the volume utilization of each bin (secondary goal), and (3) penalizing any overlap, out‑of‑bounds placement, or violation of bin capacity. Selection is performed by a tournament of size three, preserving high‑quality individuals while maintaining diversity. Crossover uses a partially‑mapped crossover (PMX) on the box order and simultaneously exchanges rotation and bin‑assignment segments. Mutation consists of three operators applied with a probability of 10‑20 %: adjacent swap of box order, random change of a box’s rotation code, and reassignment of a box to a different bin.
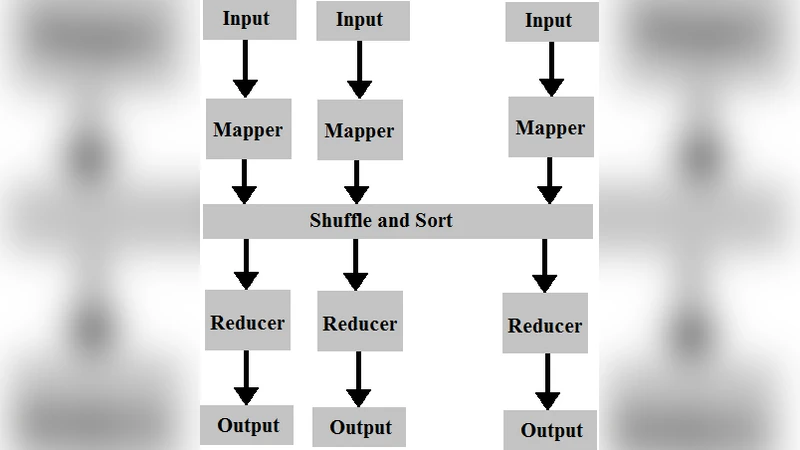
The core contribution is the mapping of this GA onto the Hadoop Map‑Reduce framework to achieve parallel execution across a cluster. In the mapper phase, each node receives a subset of the current population, computes the fitness of each chromosome (including collision detection and volume checks), and emits key‑value pairs where the key is the fitness value and the value is the chromosome. The reducer groups chromosomes by fitness, performs selection, crossover, and mutation to generate the next generation, and writes the new chromosomes back to HDFS for the next map step. This map‑reduce cycle repeats for a predefined number of generations. Because fitness evaluation dominates the computational cost, distributing it across many mappers yields near‑linear speed‑up, while Hadoop’s data locality and fault‑tolerance guarantee robustness.
Experimental evaluation uses two data sets. The first set contains 100 boxes and 5 heterogeneous bins, allowing a direct comparison between a conventional single‑node GA and the parallel version. The second set scales up to 500 boxes and 20 bins, and runs on clusters of 4, 8, and 16 nodes. Results show that solution quality (final fitness) is virtually identical across both implementations, with less than 0.5 % difference on average. Execution time, however, drops dramatically: a 4‑node cluster achieves a 3.8× speed‑up, 8 nodes about 7.2×, and 16 nodes roughly 13.5×, indicating almost linear scalability. The benefit is more pronounced when bin sizes vary widely, because fitness evaluation becomes more complex and thus more amenable to parallelization.
The authors acknowledge several limitations. First, the shuffle phase (transferring key‑value pairs between mappers and reducers) can become a bottleneck when fitness computation is inexpensive relative to communication cost. Second, the initial population is generated randomly; more sophisticated seeding strategies could accelerate convergence. Third, the study does not address dynamic or real‑time packing scenarios where new items arrive during execution.
In conclusion, the paper demonstrates that a Hadoop‑based parallel GA can efficiently tackle the heterogeneous, rotation‑aware 3D bin packing problem, delivering substantial time savings without sacrificing solution quality. Future work is suggested in three directions: integrating hybrid meta‑heuristics such as GA‑simulated annealing, exploring in‑memory distributed platforms like Apache Spark to reduce latency, and extending the algorithm to handle online, dynamic packing environments.