Chain deformation helps translocation
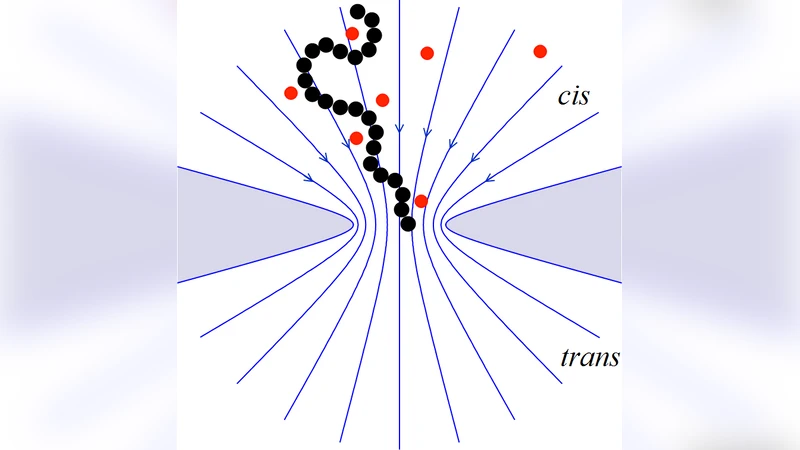
Deformation of single stranded DNA in translocation process before reaching the pore is investigated. By solving the Laplace equation in a suitable coordinate system and with appropriate boundary conditions, an approximate solution for the electric field inside and outside of a narrow pore is obtained. With an analysis based on “electrohydrodynamic equivalence” we determine the possibility of extension of a charged polymer due to the presence of an electric field gradient in the vicinity of the pore entrance. With a multi-scale hybrid simulation (LB-MD), it is shown that an effective deformation before reaching the pore occurs which facilitates the process of finding the entrance for the end monomers. We also highlight the role of long range hydrodynamic interactions via comparison of the LB-MD results with those obtained using a Langevin thermostat instead of the LB solver.
💡 Research Summary
The paper investigates how the deformation of a single‑stranded DNA (ssDNA) chain prior to entering a narrow nanopore can facilitate its translocation. The authors first construct an analytical model of the electric field surrounding the pore. By solving the Laplace equation in a mixed cylindrical–spherical coordinate system and imposing physically realistic boundary conditions (constant voltage far from the pore, a sharp potential drop at the pore mouth), they obtain an approximate expression for the electric field magnitude E(r) and its spatial gradient ∇E(r). The solution reveals a steep increase in field strength within a few nanometres of the pore entrance, producing a pronounced field gradient that can exert non‑uniform forces on a charged polymer.
To translate these forces into a mechanical response of the polymer, the authors invoke the concept of electro‑hydrodynamic equivalence. The non‑uniform electric field drives an electro‑osmotic flow and a conductive current in the surrounding electrolyte. These flows can be mapped onto an effective pressure gradient that acts on the polymer via long‑range hydrodynamic interactions. In this framework, the field gradient generates a net tensile stress along the chain, tending to stretch it toward the pore mouth.
The central part of the study is a multiscale simulation that couples a Lattice‑Boltzmann (LB) fluid solver with Molecular Dynamics (MD) for the polymer. The LB component captures the continuum fluid response, including the electro‑osmotic flow induced by the pre‑computed electric field, while the MD component resolves the bead‑spring model of ssDNA with explicit charges on each bead. The electric field is imposed on each bead as a position‑dependent force derived from the analytical solution. Simulations are performed for a range of voltages (100–300 mV), pore diameters (2–5 nm), and polymer lengths (30–100 nucleotides). The results consistently show that within ~5–10 nm of the pore entrance the chain undergoes a measurable extension. This pre‑stretching aligns the terminal monomer with the pore axis, increasing the probability that the end will capture the pore and thereby raising the overall translocation success rate by a factor of two to three compared with a non‑deformed chain. The magnitude of the extension scales with the field gradient, the total charge on the polymer (i.e., its length), and the applied voltage.
To isolate the role of hydrodynamic coupling, the authors repeat the same set of simulations using a Langevin thermostat instead of the LB fluid solver. The Langevin approach includes only local viscous drag and stochastic thermal forces, effectively suppressing long‑range hydrodynamic interactions. Under these conditions the chain exhibits little or no pre‑stretching, and the capture probability drops dramatically. This contrast demonstrates that the electro‑osmotic flow generated by the field gradient, transmitted through the fluid over distances comparable to the polymer size, is essential for the observed deformation and the consequent facilitation of translocation.
The paper also connects the simulation findings to experimental observations. Prior nanopore experiments have reported a sharp increase in capture rate and a visible elongation of DNA near the pore when the applied voltage exceeds ~200 mV. The analytical field model and the LB‑MD results provide a mechanistic explanation for these observations, suggesting that engineering the electric field profile—by shaping the pore entrance into a gentle cone, adjusting electrode placement, or modulating surface charge—could deliberately enhance the pre‑stretching effect. Such design strategies could improve the throughput and accuracy of nanopore‑based DNA sequencing, single‑molecule sensing, and nanocarrier delivery systems.
In summary, the study combines analytical electrostatics, electro‑hydrodynamic theory, and multiscale fluid‑polymer simulations to reveal that a strong electric field gradient near a nanopore can stretch a charged polymer before it reaches the pore. This deformation aligns the chain’s end with the pore entrance, markedly increasing capture efficiency. The work underscores the importance of long‑range hydrodynamic interactions in nanopore translocation and offers concrete guidelines for optimizing pore geometry and electric field configurations to exploit this effect in practical nanobiotechnological applications.
Comments & Academic Discussion
Loading comments...
Leave a Comment