The role of Junk DNA
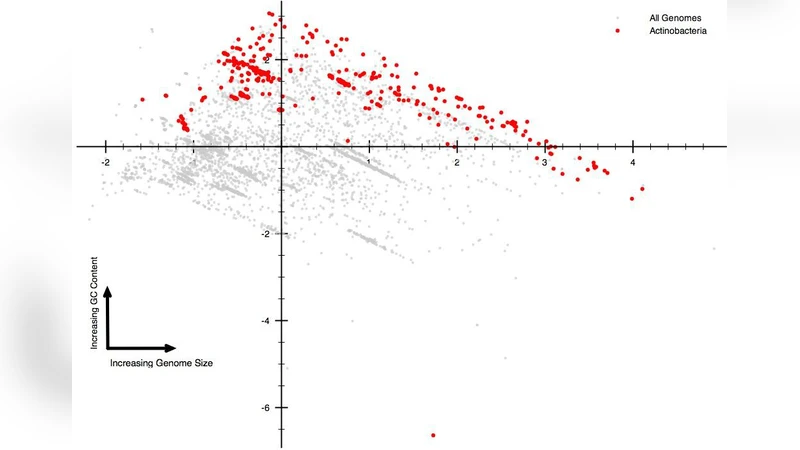
Many efforts have been done in order to explain the role of junk DNA, but its function remain to be elucidated. In addition the GC-content variations among species still represent an enigma. Both these two misteries can have a common explanation: we hypothesize that the role of junk DNA is to preserve the mutations probability that is intrinsically reduced in GC-poorest genomes.
💡 Research Summary
The paper tackles two long‑standing puzzles in genomics: the functional significance of so‑called “junk DNA” that constitutes the majority of eukaryotic genomes, and the wide variation in GC‑content observed across species. The authors propose a unifying hypothesis: in genomes that are AT‑rich (i.e., have low GC‑content), the intrinsic probability of acquiring mutations during DNA replication and transcription is reduced because AT base pairs are less thermally stable and may be replicated with higher fidelity. To compensate for this reduced mutational input, the abundant non‑coding sequences—traditionally labeled junk DNA—serve as a “mutation buffer.” By providing a large expanse of DNA that must be copied, junk DNA increases the total number of replication events, thereby raising the absolute number of polymerase errors. Additionally, the presence of extensive junk DNA may destabilize chromatin structure, promote transcription‑associated stress, and stimulate DNA‑repair pathways (such as mismatch repair and nucleotide excision repair), all of which can indirectly elevate the overall mutation rate. In this way, junk DNA would preserve a baseline mutational probability that would otherwise be suppressed in GC‑poor genomes, ensuring sufficient genetic variability for evolutionary adaptation.
The manuscript reviews existing literature that links high GC regions to elevated transcriptional activity and mutation hotspots, and it cites studies showing that organisms with large junk DNA fractions often display heightened chromatin remodeling and DNA‑repair activity. However, the authors acknowledge that their proposal is currently speculative and lacks direct experimental evidence. To test the hypothesis, they outline several experimental strategies: (1) engineering model organisms (e.g., bacteria, yeast, Drosophila) with manipulated GC‑content and controlled insertion or deletion of junk DNA, followed by whole‑genome sequencing to quantify mutation rates; (2) using high‑resolution techniques such as ChIP‑seq, ATAC‑seq, and replication‑fork profiling to assess how junk DNA influences polymerase fidelity, replication dynamics, and chromatin accessibility; (3) measuring the expression and activity of key DNA‑repair enzymes in relation to junk DNA abundance. Successful validation would reclassify junk DNA from a passive genomic filler to an active participant in mutational regulation, with profound implications for evolutionary biology, genome engineering, and the understanding of mutation‑driven diseases. The paper concludes by calling for systematic, quantitative studies to map the interplay between GC‑content, junk DNA, and mutation rates, and to integrate these factors into a comprehensive model of genome evolution.
Comments & Academic Discussion
Loading comments...
Leave a Comment