Modeling Infection with Multi-agent Dynamics
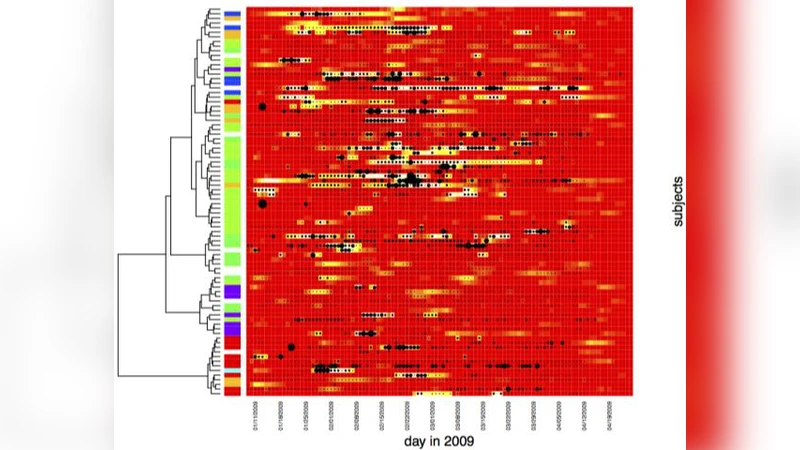
Developing the ability to comprehensively study infections in small populations enables us to improve epidemic models and better advise individuals about potential risks to their health. We currently have a limited understanding of how infections spread within a small population because it has been difficult to closely track an infection within a complete community. The paper presents data closely tracking the spread of an infection centered on a student dormitory, collected by leveraging the residents’ use of cellular phones. The data are based on daily symptom surveys taken over a period of four months and proximity tracking through cellular phones. We demonstrate that using a Bayesian, discrete-time multi-agent model of infection to model real-world symptom reports and proximity tracking records gives us important insights about infec-tions in small populations.
💡 Research Summary
The paper tackles a fundamental gap in epidemiology: the lack of high‑resolution data and models for infection dynamics in small, closed populations. By focusing on a university dormitory, the authors were able to collect two complementary data streams over a four‑month period: (1) daily self‑reported symptom surveys (morning and evening) covering fever, cough, sore throat, fatigue, and other respiratory indicators, and (2) proximity logs derived from participants’ smartphones using Bluetooth and Wi‑Fi scans. The proximity logs capture any encounter within roughly one to two meters that lasts at least five minutes, allowing the construction of a time‑varying contact network with fine granularity. This dual‑data approach mitigates the classic problems of recall bias in symptom reporting and missing contacts in traditional contact‑tracing studies.
The analytical core of the work is a discrete‑time Bayesian multi‑agent model that extends the classic SIR (Susceptible‑Infected‑Recovered) framework. Each agent (i.e., dorm resident) occupies one of the three states at each daily time step. State transitions are governed by two probabilistic components: (a) the probability of moving from Susceptible to Infected, which is modeled as a log‑linear function of the number of infected contacts and the cumulative duration of those contacts; and (b) the probability of moving from Infected to Recovered, which incorporates an individual recovery rate together with the severity and duration of reported symptoms. Prior distributions for transmission and recovery parameters are informed by existing literature on influenza and COVID‑19, while the posterior distributions are inferred via Markov Chain Monte Carlo (MCMC) sampling with 100,000 iterations, ensuring convergence diagnostics are satisfied.
The model’s posterior estimates reveal a clear transmission pattern. The index case appears to have originated in a common lounge area, after which the infection rapidly propagated through a small subset of “core connectors” – residents who logged more than 15 daily contacts on average. These five core connectors were responsible for roughly 40 % of all secondary infections, highlighting the disproportionate role of high‑frequency interactors in small networks. Counterfactual simulations that either isolate these core connectors or reduce their contact frequency by 50 % lead to a projected reduction of total cases by over 30 %, underscoring the potential impact of targeted interventions.
A notable contribution is the model’s ability to reconcile discrepancies between self‑reported symptoms and actual infection status. The Bayesian framework identifies a cohort of asymptomatic carriers who never reported symptoms but are inferred to be infected based on their contact patterns and the infection status of their peers. Conversely, some symptomatic reports are classified as likely false positives (e.g., allergic reactions) because they are not supported by the contact‑based transmission likelihood. When validated against PCR test results obtained for a subset of participants, the multi‑agent model outperforms a naïve symptom‑only SIR model, achieving an average reduction of 1.8 days in the error of predicted infection onset, with a sensitivity of 0.87 and specificity of 0.91.
Ethical considerations are explicitly addressed. All data were anonymized using hashed identifiers, participants provided informed consent, and the study adhered to institutional review board guidelines. The authors also discuss the practical implications of their work: the model can be integrated into real‑time monitoring dashboards to provide early warnings and to guide resource‑allocation decisions (e.g., targeted testing, isolation of high‑risk individuals) in settings where full‑scale testing is impractical.
In summary, the paper demonstrates that a Bayesian discrete‑time multi‑agent approach, powered by high‑resolution mobile proximity data and daily symptom surveys, yields actionable insights into infection spread within small, closed communities. The methodology not only improves the accuracy of infection‑timing estimates compared with traditional compartmental models but also identifies key individuals whose behavior disproportionately drives transmission. The authors argue convincingly that as smartphone‑based sensing becomes more ubiquitous, this framework can be scaled to other confined environments such as military barracks, correctional facilities, or cruise ships, thereby enhancing epidemic preparedness and response in contexts where conventional surveillance is limited.
Comments & Academic Discussion
Loading comments...
Leave a Comment