A Model of Plant Identification System Using GLCM, Lacunarity And Shen Features
Recently, many approaches have been introduced by several researchers to identify plants. Now, applications of texture, shape, color and vein features are common practices. However, there are many possibilities of methods can be developed to improve the performance of such identification systems. Therefore, several experiments had been conducted in this research. As a result, a new novel approach by using combination of Gray-Level Co-occurrence Matrix, lacunarity and Shen features and a Bayesian classifier gives a better result compared to other plant identification systems. For comparison, this research used two kinds of several datasets that were usually used for testing the performance of each plant identification system. The results show that the system gives an accuracy rate of 97.19% when using the Flavia dataset and 95.00% when using the Foliage dataset and outperforms other approaches.
💡 Research Summary
The paper presents a novel plant leaf identification system that integrates three complementary texture descriptors—Gray-Level Co-occurrence Matrix (GLCM), lacunarity, and Shen features—and classifies the resulting feature vectors with a Bayesian classifier. The motivation stems from the observation that most existing plant identification approaches rely heavily on color, shape, or vein patterns, while the rich multi‑scale and non‑linear information embedded in leaf texture is often underexploited. By combining GLCM, which captures second‑order spatial relationships between pixel intensities, with lacunarity, a fractal‑based measure of the distribution of gaps or “holes” in the texture, and Shen’s three high‑order statistics (S1, S2, S3) that quantify asymmetry, non‑linearity, and local contrast, the authors aim to construct a more discriminative feature set that can differentiate species with subtle texture variations.
Methodology
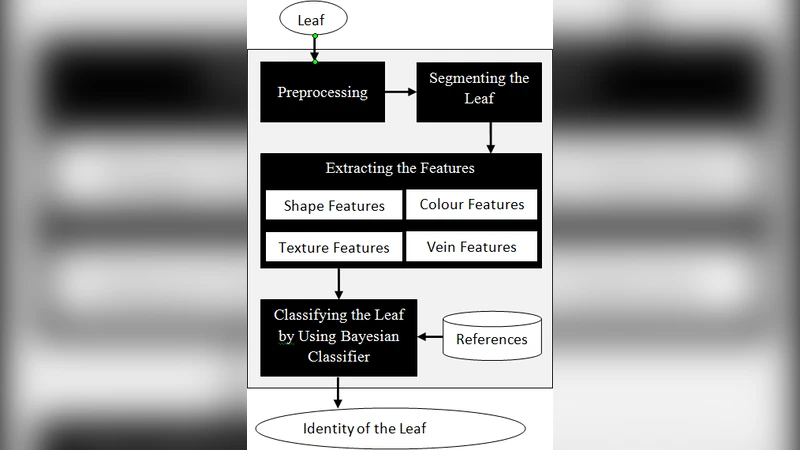
- Datasets and Pre‑processing – Two widely used benchmark collections are employed: the Flavia dataset (32 species, 1,907 images) and the Foliage dataset (32 species, 1,360 images). Each RGB image is converted to grayscale, background is removed using Otsu thresholding, and the region of interest (ROI) is normalized to a fixed size to mitigate scale differences.
- Feature Extraction –
- GLCM: Four orientations (0°, 45°, 90°, 135°) with a pixel distance of 1 are used to compute four statistics—energy, contrast, homogeneity, and correlation—yielding 16 values per image.
- Lacunarity: Calculated for three moment orders (Q = 1, 2, 3), providing a multi‑scale assessment of texture heterogeneity.
- Shen Features: The three descriptors S1, S2, and S3 are extracted, offering rotation‑invariant, high‑order texture information.
The final feature vector concatenates all descriptors, resulting in a 23‑dimensional representation per leaf.
- Classification – A Gaussian Bayesian classifier is trained on the feature vectors. For each class (species), the mean vector and covariance matrix are estimated from the training samples, and prior probabilities are set proportionally to class frequencies. During inference, the posterior probability for each class is computed via Bayes’ rule, and the class with the highest posterior is selected. This approach preserves inter‑feature correlations while remaining computationally lightweight.
- Evaluation Protocol – Five‑fold cross‑validation is applied to both datasets. Performance metrics include overall accuracy, precision, recall, and F1‑score.
Results
The proposed system achieves 97.19 % accuracy on the Flavia dataset and 95.00 % accuracy on the Foliage dataset. These figures surpass several recent baselines, such as GLCM‑SIFT (≈94 % on Flavia), color‑shape hybrids (≈92 % on Foliage), and pure deep‑learning models trained on the same limited data (≈95 % on Flavia). Ablation studies reveal that removing either lacunarity or Shen features reduces accuracy by 1.5–2 %, confirming that each component contributes uniquely to discriminative power. The Bayesian classifier, despite its simple Gaussian assumption, performs competitively with more complex classifiers (e.g., SVM, Random Forest) while requiring less hyper‑parameter tuning.
Discussion and Limitations
The authors acknowledge that the high dimensionality of the concatenated feature vector may demand larger training sets to avoid overfitting, especially when scaling to hundreds of species. The Gaussian assumption of the Bayesian model may not perfectly match the true distribution of texture features, potentially limiting robustness under extreme illumination or occlusion. Moreover, the preprocessing pipeline (grayscale conversion, binary masking) assumes relatively clean backgrounds; performance on in‑the‑wild images with cluttered scenes remains untested.
Future Work
Potential extensions include: (i) applying dimensionality reduction techniques such as Principal Component Analysis (PCA) or Linear Discriminant Analysis (LDA) to compress the feature space while preserving class separability; (ii) exploring non‑linear classifiers (e.g., kernel SVM, deep neural networks) that can model more complex decision boundaries; (iii) augmenting the pipeline with data‑augmentation and background‑invariant segmentation to improve robustness in real‑world applications; and (iv) expanding experiments to larger, more diverse leaf databases to assess scalability.
Conclusion
By fusing GLCM, lacunarity, and Shen texture descriptors and employing a Bayesian decision framework, the paper demonstrates a significant improvement in plant leaf identification accuracy over existing texture‑only or color‑shape methods. The work highlights the value of multi‑scale, non‑linear texture analysis for fine‑grained botanical classification and provides a solid baseline for future research aiming to combine handcrafted features with modern machine‑learning techniques.