Stochastic synchronization of genetic oscillators induced by miRNA
We study two important roles of miRNA as stress inducer and synchronizing agent in regulating diffusively coupled genetic oscillators within stochastic formalism and competition between them. We identify optimal value of coupling constant at which the rate of synchronization is strongest, below this value sychronizing activity dominates stress activity, and above it stress activity destroys synchronizing pattern. The concentration of miRNA maintained in individual oscillator modulates the synchronization activity achieved by diffusing miRNA.
💡 Research Summary
In this study the authors investigate how micro‑RNA (miRNA) can simultaneously act as a synchronizing messenger and a stress‑inducing agent in a pair of diffusively coupled genetic oscillators. Starting from the well‑known Vilar model of a circadian‑like transcription‑translation feedback loop, they extend the network to include two genes (an activator and a repressor), their mRNAs, the corresponding proteins, and the formation of an activator‑repressor complex. miRNA is introduced as a separate molecular species that can bind the repressor mRNA to form a C_RISC complex, thereby repressing translation. The kinetic scheme comprises 22 molecular species and 21 reactions, each with explicit rate constants taken from previous literature or estimated for the miRNA processes (synthesis k19, degradation k20, complex formation k21, and complex degradation k22).
Two identical oscillators are coupled through the diffusion of miRNA between them. The coupling is represented by two reversible reactions with rate constants ε and ε′ (set equal in most simulations). The authors employ Gillespie’s stochastic simulation algorithm (SSA) to generate time courses for the activator (A) and repressor (R) protein concentrations in each oscillator, as well as two‑dimensional recurrence plots to visualize correlation.
Key findings:
-
Zero coupling (ε = 0) – The oscillators behave independently; recurrence plots show a random cloud of points, confirming no synchrony.
-
Moderate coupling (ε ≈ 3) – Both A and R trajectories become highly correlated; points in the recurrence plots concentrate along the diagonal, indicating strong stochastic synchrony despite intrinsic noise. This value is identified as the optimal coupling strength for maximal synchronization rate.
-
Strong coupling (ε ≥ 15) – Correlation collapses; the recurrence plots become scattered again. The authors interpret this as miRNA‑mediated stress overwhelming the synchronizing influence, effectively disrupting the feedback loops that generate the oscillations.
-
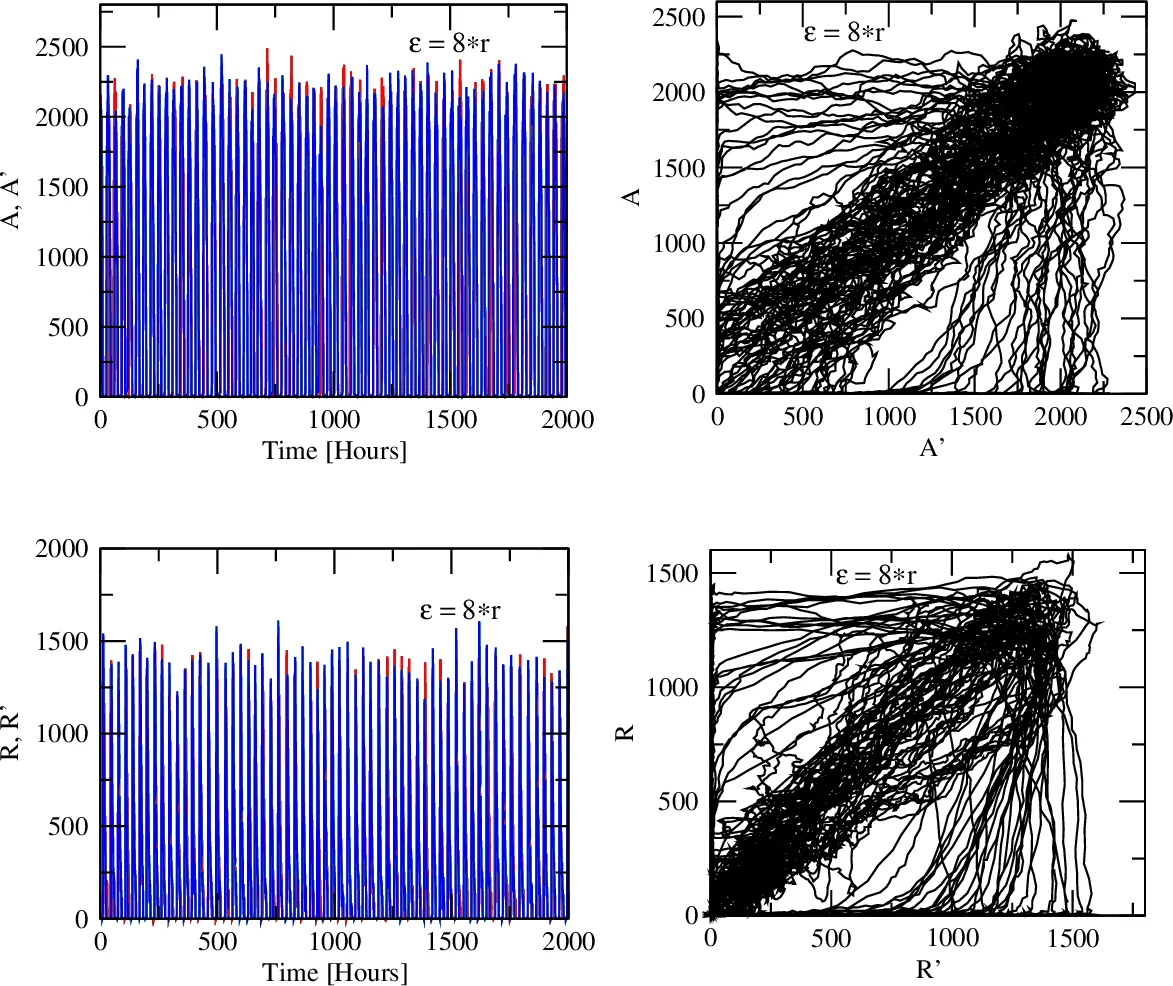
Random coupling (ε = C·r, where r is a random number) – Synchronization can still be achieved but requires a larger effective coupling magnitude (C = 8). This demonstrates that variability in miRNA diffusion demands higher average coupling to maintain synchrony.
-
Effect of miRNA synthesis and degradation – Varying the synthesis rate k19 (values 18–22) shifts the optimal ε upward, but excessively high k19 destroys the oscillatory behavior altogether. The degradation rate k20 shows a non‑monotonic influence: ε first increases with k20, reaches a peak, then declines, suggesting an optimal miRNA turnover for synchronization.
-
Complex formation and decay – Increasing the complex formation rate k21 raises ε, while increasing the complex decay rate k22 lowers ε. Thus, the balance between miRNA‑mRNA binding and complex turnover directly modulates the coupling efficiency.
Overall, the paper demonstrates a competition between two miRNA‑driven processes: (i) diffusion‑mediated synchronization of genetic clocks, and (ii) stress induction through excessive repression of the repressor gene. The system exhibits a bell‑shaped dependence on coupling strength and miRNA concentration, with an optimal window where synchrony is strongest. The authors argue that such a mechanism could be relevant for circadian rhythm regulation, where circulating miRNAs have been observed in body fluids, and for pathological conditions where miRNA levels are dysregulated (e.g., cancer, metabolic and inflammatory diseases). By highlighting the dual role of miRNA, the work suggests new avenues for therapeutic manipulation of cellular clocks and for using miRNA as a biomarker of stress‑related dysregulation.
Comments & Academic Discussion
Loading comments...
Leave a Comment