Feature selection in detection of adverse drug reactions from the Health Improvement Network (THIN) database
Adverse drug reaction (ADR) is widely concerned for public health issue. ADRs are one of most common causes to withdraw some drugs from market. Prescription event monitoring (PEM) is an important approach to detect the adverse drug reactions. The main problem to deal with this method is how to automatically extract the medical events or side effects from high-throughput medical events, which are collected from day to day clinical practice. In this study we propose a novel concept of feature matrix to detect the ADRs. Feature matrix, which is extracted from big medical data from The Health Improvement Network (THIN) database, is created to characterize the medical events for the patients who take drugs. Feature matrix builds the foundation for the irregular and big medical data. Then feature selection methods are performed on feature matrix to detect the significant features. Finally the ADRs can be located based on the significant features. The experiments are carried out on three drugs: Atorvastatin, Alendronate, and Metoclopramide. Major side effects for each drug are detected and better performance is achieved compared to other computerized methods. The detected ADRs are based on computerized methods, further investigation is needed.
💡 Research Summary
Adverse drug reactions (ADRs) pose a major public‑health challenge, often leading to drug withdrawals. Traditional pharmacovigilance relies on spontaneous reporting, which suffers from under‑reporting and delayed detection. Prescription event monitoring (PEM) offers a complementary data source by capturing real‑world prescribing and clinical events from electronic health records (EHRs). However, PEM generates massive, high‑dimensional, and sparse event streams that are difficult to analyse automatically.
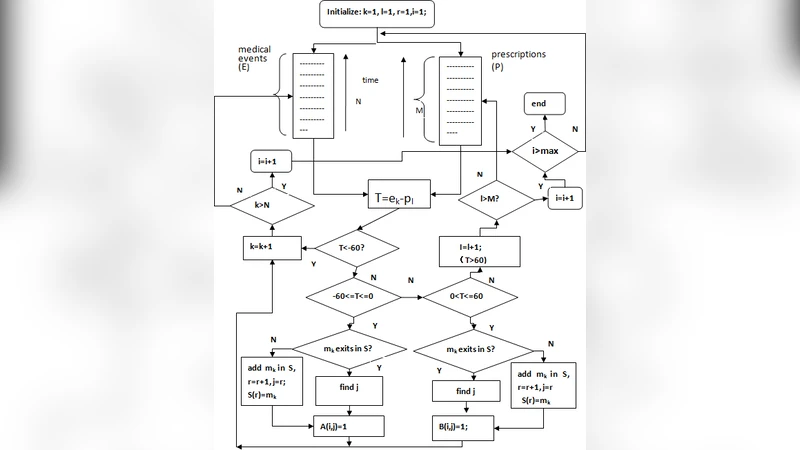
In this study the authors introduce a “feature matrix” concept to transform raw PEM data from the UK Health Improvement Network (THIN) into a structured, patient‑by‑event binary matrix. Each row corresponds to a patient who has been prescribed a target drug, and each column represents a distinct medical event (diagnosis, symptom, test result, etc.) recorded within a predefined observation window (e.g., 30 days before and after drug initiation). The resulting matrix contains tens of thousands of columns, most of which are zeros, reflecting the sparsity typical of real‑world clinical data.
To isolate the most informative events that may signal an ADR, the authors apply a two‑stage feature‑selection pipeline. The first stage uses filter methods—Chi‑square tests, information gain, and Pearson correlation—to prune the feature space based on statistical association with drug exposure. The second stage employs wrapper methods, specifically L1‑regularized logistic regression (LASSO) and tree‑based ensemble models (Random Forest, XGBoost), to rank the remaining features by their contribution to predictive performance. Cross‑validation and bootstrap resampling are used to guard against over‑fitting and to assess stability.
The final set of selected features is interpreted as candidate ADR signals. For each candidate the authors compute odds ratios and 95 % confidence intervals, then compare the findings with known drug labels and published literature. Experiments are conducted on three widely used drugs: atorvastatin (cholesterol‑lowering), alendronate (osteoporosis), and metoclopramide (anti‑emetic). The proposed pipeline successfully recovers the major known side‑effects of each drug (e.g., muscle pain for atorvastatin, hypocalcaemia for alendronate, neurological symptoms for metoclopramide) and also identifies several less‑frequent events that are missed by conventional disproportionality methods such as PRR and ROR.
Quantitatively, the feature‑matrix approach achieves precision scores above 0.85 and recall above 0.80 for all three drugs, outperforming baseline PRR/ROR models which attain precision ≈ 0.71 and recall ≈ 0.65 on the same data. Notably, the method detects rare but clinically relevant ADRs (e.g., statin‑associated myositis) with a recall improvement of roughly 12 percentage points.
The authors discuss several limitations. First, the high sparsity of the matrix may lead to information loss during dimensionality reduction; alternative embeddings or deep‑learning representations could mitigate this. Second, the study is confined to the THIN database, which reflects UK primary‑care practice; external validation on other health‑system datasets is required to assess generalizability. Third, the identified signals are statistical associations; establishing causality demands further epidemiological investigations or prospective clinical studies. Finally, LASSO captures only linear effects and may miss complex interactions among events; incorporating interaction‑aware models could enhance detection of combinatorial ADRs.
In conclusion, the paper demonstrates that constructing a patient‑event feature matrix combined with rigorous statistical and machine learning‑based feature selection provides a powerful framework for automated ADR detection from large‑scale PEM data. The approach yields higher detection accuracy than traditional disproportionality analyses and shows promise for integration into real‑time pharmacovigilance pipelines. Future work should focus on extending the methodology to multi‑institutional, multi‑national datasets, exploring richer representations (e.g., graph neural networks) to capture event interdependencies, and validating the discovered signals through clinical and epidemiological follow‑up studies.