Computer simulations reveal complex distribution of haemodynamic forces in a mouse retina model of angiogenesis
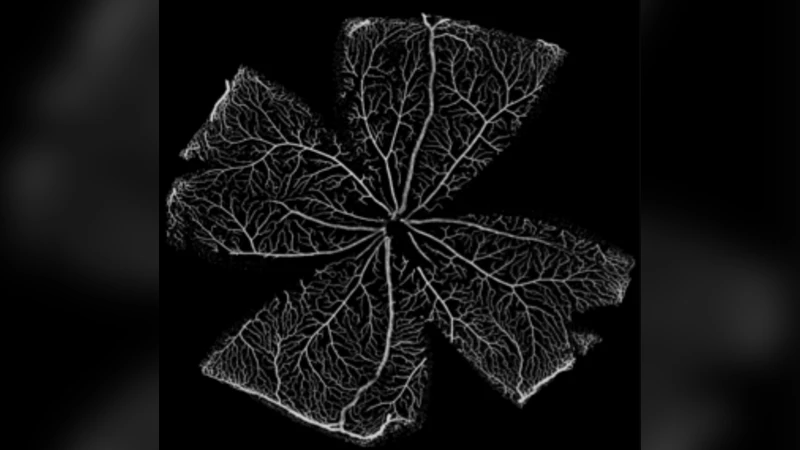
There is currently limited understanding of the role played by haemodynamic forces on the processes governing vascular development. One of many obstacles to be overcome is being able to measure those forces, at the required resolution level, on vessels only a few micrometres thick. In the current paper, we present an in silico method for the computation of the haemodynamic forces experienced by murine retinal vasculature (a widely used vascular development animal model) beyond what is measurable experimentally. Our results show that it is possible to reconstruct high-resolution three-dimensional geometrical models directly from samples of retinal vasculature and that the lattice-Boltzmann algorithm can be used to obtain accurate estimates of the haemodynamics in these domains. We generate flow models from samples obtained at postnatal days (P) 5 and 6. Our simulations show important differences between the flow patterns recovered in both cases, including observations of regression occurring in areas where wall shear stress gradients exist. We propose two possible mechanisms to account for the observed increase in velocity and wall shear stress between P5 and P6: i) the measured reduction in typical vessel diameter between both time points, ii) the reduction in network density triggered by the pruning process. The methodology developed herein is applicable to other biomedical domains where microvasculature can be imaged but experimental flow measurements are unavailable or difficult to obtain.
💡 Research Summary
**
This paper addresses the challenge of quantifying haemodynamic forces in the developing mouse retina, a widely used model for studying vascular development, where experimental measurement of flow at micrometre resolution is practically impossible. The authors present a fully integrated in‑silico workflow that starts from high‑resolution confocal microscopy images of retinal vasculature at post‑natal days 5 (P5) and 6 (P6), proceeds through automated image segmentation, skeletonisation, and three‑dimensional lumen reconstruction, and culminates in lattice‑Boltzmann method (LBM) simulations of blood flow using the open‑source CFD solver HemeLB.
In the imaging stage, raw fluorescence stacks are denoised, thresholded, and binarised to obtain a vessel lumen mask. A multi‑scale morphological pipeline extracts centrelines and local radii, preserving branch points and loops. These geometric descriptors are then swept into a surface mesh with a resolution finer than one‑tenth of the smallest vessel diameter (≈0.5 µm), ensuring that geometric errors do not dominate the fluid dynamics calculations.
The fluid dynamics stage treats blood as a non‑Newtonian fluid, employing a Carreau‑Yasuda viscosity model calibrated from literature data to capture shear‑thinning behaviour at low shear rates (≈600 s⁻¹). Boundary conditions are set using physiological values: mean arterial pressure (≈80 mmHg) at the arterial inlet, intra‑ocular pressure (≈15 mmHg) at the venous outlet, yielding a driving pressure of about 65 mmHg. Inlet velocity is initialized from experimentally reported mean arterial speeds (≈25 mm s⁻¹), while the outlet uses a free‑pressure (Neumann) condition. The LBM discretisation maps the reconstructed geometry onto a regular lattice, allowing massive parallelisation on GPU/CPU clusters and rapid convergence even in highly branched networks.
Simulation results reveal that between P5 and P6 the average vessel diameter contracts by roughly 15 % (from ~7 µm to ~6 µm) and the network connectivity density drops by about 20 %. These structural changes lead to a 30 % increase in mean flow velocity and a comparable rise in wall shear stress (WSS). Importantly, regions with steep WSS gradients—typically near bifurcations—coincide with observed vessel regression (pruning), suggesting that mechanical cues may trigger endothelial cell apoptosis or remodeling. The authors propose two complementary mechanisms for the observed haemodynamic escalation: (i) reduced lumen diameter lowers hydraulic resistance, amplifying velocity and WSS under the same pressure drop; (ii) pruning reduces the number of parallel pathways, concentrating flow through the remaining vessels and further elevating local shear.
Method validation is performed on a simple cylindrical tube where analytical Poiseuille solutions are available; the LBM predictions deviate by less than 2 % in both velocity and pressure loss. Comparisons with traditional one‑dimensional network models demonstrate that the three‑dimensional LBM captures complex flow features such as recirculation zones and non‑uniform shear distributions that 1‑D approaches miss.
The paper’s contributions are threefold: (1) an open‑source, reproducible pipeline for converting high‑resolution vascular images into CFD‑ready geometries; (2) demonstration that LBM can reliably resolve flow in vessels as small as 5 µm, providing quantitative estimates of velocity and WSS where experimental techniques fail; (3) mechanistic insight into how morphological remodeling (diameter reduction and network pruning) drives haemodynamic changes during early retinal angiogenesis.
Limitations include the neglect of red‑blood‑cell aggregation effects (Fåhræus‑Lindqvist) and vessel wall elasticity, as well as the use of global pressure boundary conditions that do not capture local pressure heterogeneities. Future work is outlined to incorporate fluid‑structure interaction, multi‑phase blood models, and spatially resolved pressure measurements, which would enhance predictive fidelity. The authors anticipate that the workflow will be transferable to other microvascular systems—brain, tumor, or peripheral tissues—where direct haemodynamic measurement is equally challenging, thereby providing a valuable computational tool for vascular biology and translational research.
Comments & Academic Discussion
Loading comments...
Leave a Comment