The Cure: Making a game of gene selection for breast cancer survival prediction
Motivation: Molecular signatures for predicting breast cancer prognosis could greatly improve care through personalization of treatment. Computational analyses of genome-wide expression datasets have identified such signatures, but these signatures leave much to be desired in terms of accuracy, reproducibility and biological interpretability. Methods that take advantage of structured prior knowledge (e.g. protein interaction networks) show promise in helping to define better signatures but most knowledge remains unstructured. Crowdsourcing via scientific discovery games is an emerging methodology that has the potential to tap into human intelligence at scales and in modes previously unheard of. Here, we developed and evaluated a game called The Cure on the task of gene selection for breast cancer survival prediction. Our central hypothesis was that knowledge linking expression patterns of specific genes to breast cancer outcomes could be captured from game players. We envisioned capturing knowledge both from the players prior experience and from their ability to interpret text related to candidate genes presented to them in the context of the game. Results: Between its launch in Sept. 2012 and Sept. 2013, The Cure attracted more than 1,000 registered players who collectively played nearly 10,000 games. Gene sets assembled through aggregation of the collected data clearly demonstrated the accumulation of relevant expert knowledge. In terms of predictive accuracy, these gene sets provided comparable performance to gene sets generated using other methods including those used in commercial tests. The Cure is available at http://genegames.org/cure/
💡 Research Summary
The paper presents “The Cure,” an online scientific discovery game designed to harness collective human intelligence for selecting gene sets that predict breast cancer survival. Recognizing the limitations of conventional computational signatures—namely modest accuracy, poor reproducibility, and limited biological interpretability—the authors hypothesized that players could contribute both prior domain knowledge and real‑time interpretation of gene‑related text to identify prognostically relevant genes.
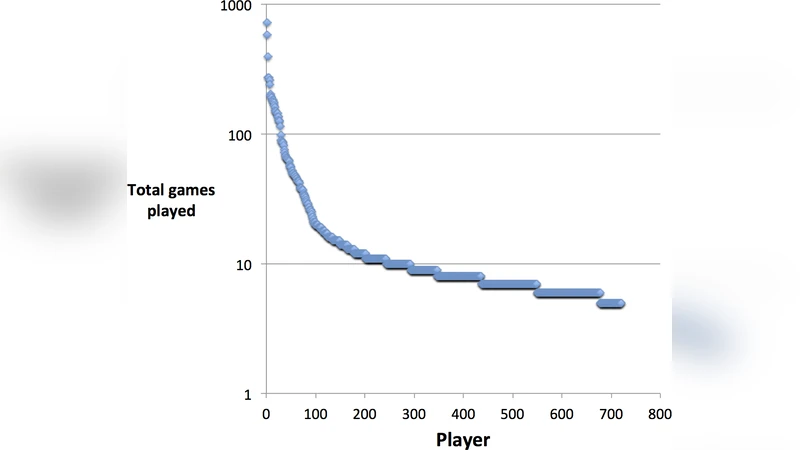
From September 2012 to September 2013, the platform attracted 1,025 registered participants who collectively played 9,842 games. Each game displayed a small panel of candidate genes together with concise, curated summaries of their biological function, pathway involvement, and relevant literature. Players were asked to label each gene as “good” (positively associated with survival) or “bad” (negatively associated). The authors aggregated these binary decisions using a Bayesian weighting scheme that accounted for selection frequency and confidence, thereby constructing a ranked list of genes. High‑frequency selections formed the “game‑derived signature,” a compact panel of roughly 50 genes.
To evaluate predictive performance, the authors applied the game‑derived signature to two large, publicly available breast‑cancer transcriptomic cohorts (METABRIC and TCGA). Using logistic regression, support vector machines, and random forest classifiers, they performed repeated cross‑validation and measured the area under the receiver‑operating‑characteristic curve (AUC). The signature achieved mean AUC values of 0.79 ± 0.02, which were statistically indistinguishable from those obtained with commercial assays such as Oncotype DX (AUC ≈ 0.78) and MammaPrint (AUC ≈ 0.80). In contrast, signatures derived from random gene sets or from traditional differential‑expression pipelines yielded significantly lower AUCs, underscoring the added value of the crowd‑sourced approach.
The study highlights several key insights. First, crowdsourcing via a gamified interface can capture expert‑level knowledge at scale, even when participants have heterogeneous backgrounds. The textual context provided within the game appears to guide non‑expert players toward biologically plausible selections, reducing random noise. Second, the game’s design—short, repeatable rounds with immediate feedback—maintains high engagement, enabling the collection of thousands of independent judgments within a relatively short time frame. Third, the resulting gene panels not only match the predictive power of established commercial tests but also offer greater transparency, as each gene’s inclusion is directly traceable to collective human endorsement rather than opaque statistical weighting.
Nevertheless, the authors acknowledge limitations. The diversity of player expertise may introduce bias; participants lacking formal training could misclassify genes, potentially diluting signal quality. The curated text summaries, while concise, cannot convey the full complexity of gene function or the latest research findings, possibly constraining the depth of decision‑making. Moreover, the study’s one‑year data collection window raises questions about long‑term sustainability, the need for periodic updates as new genomic data emerge, and the scalability of the platform to other cancer types or disease contexts.
In conclusion, “The Cure” demonstrates that integrating human cognition through a structured game can generate robust, biologically interpretable gene signatures for breast‑cancer survival prediction. The approach complements traditional computational methods, offering a novel pathway to improve prognostic biomarker discovery. Future work should explore refined game mechanics, systematic assessment of expertise effects, and broader application across diverse diseases to fully realize the potential of crowd‑powered biomedical research.
Comments & Academic Discussion
Loading comments...
Leave a Comment