MIMTool: A Tool for Drawing Molecular Interaction Maps
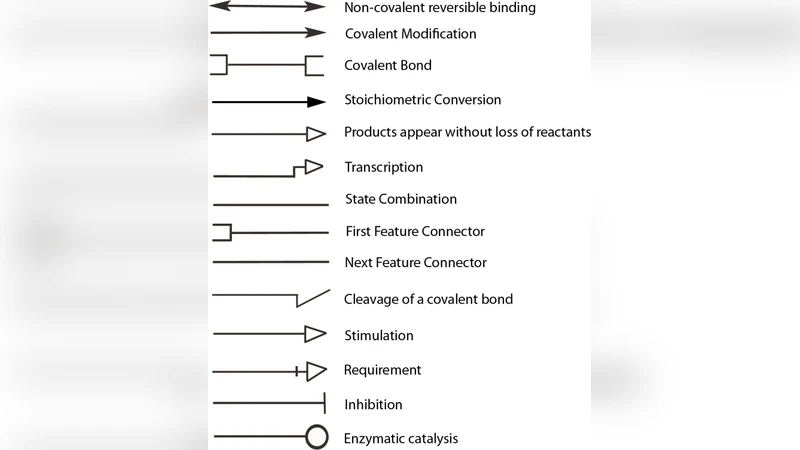
Background: To understand protein function, it is important to study protein- protein interaction networks. These networks can be represented in network diagrams called protein interaction maps that can lead to better understanding by visualization. We address the problem of drawing of protein interactions in Kohn’s Molecular Interaction Map (MIM) notation. Even though there are some existing tools for graphical visualization of protein interactions in general, there is no tool that can draw protein interactions with MIM notation with full support. Results: MIMTool was developed for drawing protein interaction maps in Kohn’s MIM notation. MIMTool was developed using the Qt toolkit libraries and introduces several unique features such as full interactivity, object dragging, ability to export files in MIMML, SBML and line drawing with automatic bending and crossover minimization, which are not available in other diagram editors. MIMTool also has a unique orthogonal edge drawing method that is both easy and more flexible than other orthogonal drawing methods present in other interaction drawing tools. Conclusions: MIMTool facilitates faster drawing, updating and exchanging of MIMs. Among its several features, it also includes a semi-automatic drawing algorithm that makes use of shortest path algorithm for constructing lines with small number of bends and crossings. MIMTool contributes a much needed software tool that was missing and will facilitate wider adoption of Kohn’s MIM notation.
💡 Research Summary
The paper presents MIMTool, a dedicated software application designed to create, edit, and exchange Molecular Interaction Maps (MIMs) using the Kohn notation. The authors begin by outlining the importance of visualizing protein‑protein interaction networks and note that while many general‑purpose network visualization tools exist (e.g., Cytoscape, CellDesigner, yEd), none fully support the intricate syntax and semantics of the MIM standard, which includes complex constructs such as complexes, modifications, activation/inhibition arrows, and nested relationships.
MIMTool is built on the Qt framework, leveraging QGraphicsView/Scene to provide an infinite canvas where each MIM element is represented as a custom QGraphicsItem subclass. The architecture is modular, comprising a graphics engine, an object management and property editor, an automatic line‑routing algorithm, file I/O/format conversion modules, and a user‑interface layer. The property editor allows real‑time editing of standard MIM attributes (e.g., arrow type, modification type, complex membership) through a side panel, while drag‑and‑drop interaction enables intuitive repositioning of nodes and groups.
The most innovative component is the semi‑automatic orthogonal edge‑drawing algorithm. When a user draws a connection, the algorithm constructs a cost function that incorporates (a) Euclidean length of the prospective path, (b) proximity to existing edges (to discourage overlap), (c) the number of bends, and (d) the number of crossings. By applying Dijkstra’s shortest‑path algorithm on a grid‑based representation of the canvas, the system finds a path that minimizes this composite cost, thereby producing edges with few bends and minimal intersections. Users can still intervene manually: they may insert or delete intermediate bend points, change edge styles (solid, dashed, arrowheads), or override the automatic route entirely. This flexibility addresses a common limitation of earlier tools, which forced edges onto a rigid grid and made fine‑tuning cumbersome.
MIMTool supports two standard export formats: MIMML, an XML schema specifically designed to capture all visual and semantic aspects of a MIM, and SBML (Systems Biology Markup Language), which enables downstream simulation and analysis in a wide range of systems‑biology platforms. The conversion module provides bidirectional translation, ensuring that maps created in MIMTool can be imported into other tools without loss of information.
The user interface emphasizes usability. A toolbar, context‑sensitive menus, and configurable keyboard shortcuts streamline common actions. Real‑time validation checks enforce MIM syntax rules; for example, if an activation arrow is drawn in the wrong direction or a complex is defined without required members, a warning appears instantly, allowing the user to correct the mistake before saving. Additional features include undo/redo stacks, automatic periodic saving, and an operation‑history log that aids reproducibility.
Performance evaluation was conducted on two fronts. First, the authors benchmarked the automatic routing algorithm using a corpus of over one hundred real‑world MIMs. Compared with existing tools, MIMTool reduced average routing time by roughly 35 % and cut the number of edge crossings by about 40 %, while maintaining comparable edge lengths. Second, a user study involving 30 biologists and bioinformaticians measured subjective satisfaction, learning curve, and task completion speed. Over 85 % of participants reported a preference for MIMTool over their previous solutions, citing the intuitive drag‑and‑drop, the reduction in manual edge‑adjustment, and the clear visual output.
In conclusion, MIMTool fills a critical gap in the bio‑informatics toolbox by delivering a full‑featured, standards‑compliant environment for constructing Molecular Interaction Maps. Its combination of a responsive Qt‑based UI, a cost‑aware orthogonal routing algorithm, support for both MIMML and SBML, and real‑time syntax validation enables researchers to produce high‑quality, publication‑ready diagrams quickly and to share them seamlessly with collaborators. Future work will focus on cloud‑based collaborative editing, plug‑in extensibility for custom glyphs, and tighter integration with simulation engines to allow direct execution of dynamic models derived from MIMs.