Density Adaptive Parallel Clustering
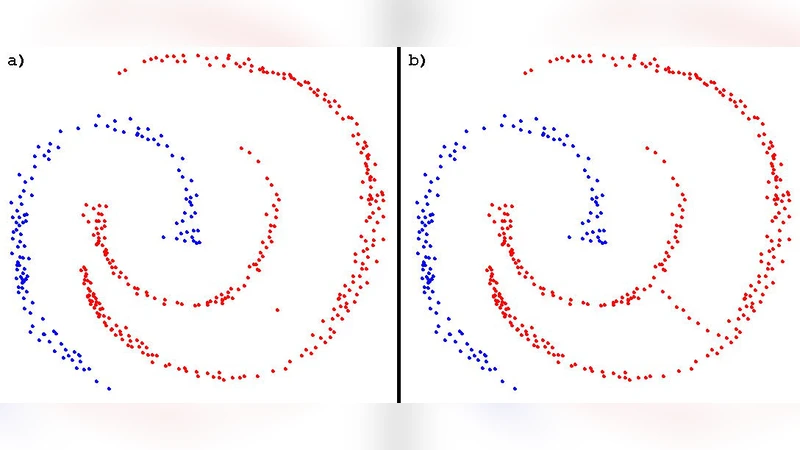
In this paper we are going to introduce a new nearest neighbours based approach to clustering, and compare it with previous solutions; the resulting algorithm, which takes inspiration from both DBscan and minimum spanning tree approaches, is deterministic but proves simpler, faster and doesnt require to set in advance a value for k, the number of clusters.
💡 Research Summary
**
The paper proposes a novel density‑adaptive, parallel clustering algorithm that seeks to combine the strengths of DBSCAN and minimum‑spanning‑tree (MST) approaches while eliminating the need for a globally fixed number of clusters or a single set of density parameters. The authors observe that naïve nearest‑neighbour merging (using a fixed number m of nearest neighbours) quickly fragments data and suffers from the classic “single‑link effect,” where points that are roughly equidistant to two dense regions cause unintended merges. To overcome this, they incorporate the concept of density‑reachability from DBSCAN, but instead of using a global radius ε and a global minimum‑neighbour count m, they infer these values locally for each point, allowing the algorithm to adapt to regions of varying density.
The method proceeds in three main stages:
-
Pre‑clustering (Canopy) – The entire dataset is first partitioned using canopy clustering, a fast O(n) heuristic that creates overlapping “pre‑clusters” based on two distance thresholds (T₁ > T₂). This step provides a coarse spatial decomposition that can be processed independently.
-
Map (local processing) – For each canopy region, an SS+‑tree (a spatial split‑tree variant) is built. The tree enables logarithmic‑time range queries. For every point p in the region, the algorithm:
- Determines a local radius εₚ (the paper suggests using statistics of the k‑nearest‑neighbour distances within the region, though the exact formula is not specified).
- Retrieves all points within εₚ using the SS+‑tree (average cost O(log k) per query, where k is the number of points in the region).
- If at least m neighbours are found, p and all neighbours are merged into the same cluster using a Union‑Find data structure with union‑by‑rank and path compression. The average cost of each merge is O(m α(k)), where α is the inverse Ackermann function (practically constant).
-
Reduce (global merging) – Because canopy regions overlap, points on region boundaries may belong to multiple local Union‑Find structures. The reduce step merges these structures, again using Union‑Find, to obtain a globally consistent labeling.
The authors argue that this pipeline yields near‑linear overall complexity: canopy clustering O(n), SS+‑tree construction O(k log k) per region, and per‑point processing O(log k + m α(k)). Assuming m and the average number of points returned by a range query are bounded by a small constant, the total expected runtime is O(n log n). Memory usage is also modest because each region is processed independently, allowing the algorithm to scale to datasets that do not fit entirely in RAM.
Key contributions highlighted by the paper include:
- Parameter‑free density adaptation – By estimating ε locally, the algorithm can handle heterogeneous density without manual tuning, a known limitation of classic DBSCAN.
- Deterministic outcome – Unlike many hierarchical or stochastic methods, the use of deterministic data structures (SS+‑tree, Union‑Find) guarantees reproducible results.
- High parallelizability – The canopy‑based decomposition naturally maps to a Map‑Reduce framework; each region can be assigned to a separate processor or node, and the final reduction step is embarrassingly parallel.
However, the paper has several notable shortcomings:
- Lack of concrete ε estimation formula – The method for deriving a per‑point radius is described only qualitatively. Without a precise algorithm, reproducibility and implementation become ambiguous.
- Residual dependence on user‑defined parameters – The smoothing parameter m and the canopy thresholds (T₁, T₂) still require user selection. Although the authors claim reduced sensitivity, no systematic guidance or sensitivity analysis is provided.
- Missing empirical evaluation – The manuscript does not include experiments on synthetic or real‑world datasets, nor does it report runtime, memory consumption, or clustering quality (e.g., Adjusted Rand Index, silhouette scores) compared to DBSCAN, OPTICS, or MST‑based methods. Consequently, the claimed performance gains remain speculative.
- Scalability of SS+‑trees in high dimensions – While the authors argue that SS+‑trees scale better than R‑trees, they do not discuss the curse of dimensionality, which can degrade range‑query performance dramatically. No dimensionality experiments are presented.
- Potential persistence of single‑link effect – Although local ε estimation mitigates accidental merges, the algorithm still merges any point that has m neighbours within εₚ, regardless of whether those neighbours belong to distinct dense regions. In datasets with long chains of sparse points, the single‑link phenomenon may reappear.
In summary, the paper introduces an interesting hybrid clustering framework that blends density‑based reasoning with efficient spatial indexing and a parallel Map‑Reduce architecture. The theoretical analysis suggests near‑linear time complexity and deterministic behavior, and the idea of locally adaptive density thresholds addresses a well‑known weakness of DBSCAN. Nonetheless, the absence of concrete parameter‑estimation procedures, a thorough experimental validation, and a discussion of high‑dimensional behavior limits the immediate impact of the work. Future research should focus on formalizing the ε‑estimation step, providing extensive benchmarks on large, high‑dimensional datasets, and exploring adaptive strategies for the remaining user‑controlled parameters to fully realize the algorithm’s potential in practical big‑data clustering scenarios.
Comments & Academic Discussion
Loading comments...
Leave a Comment