Visualization and Correction of Automated Segmentation, Tracking and Lineaging from 5-D Stem Cell Image Sequences
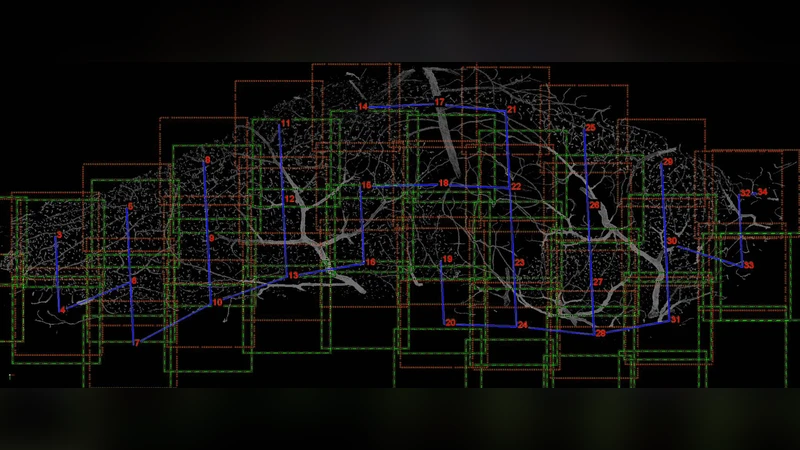
Results: We present an application that enables the quantitative analysis of multichannel 5-D (x, y, z, t, channel) and large montage confocal fluorescence microscopy images. The image sequences show stem cells together with blood vessels, enabling quantification of the dynamic behaviors of stem cells in relation to their vascular niche, with applications in developmental and cancer biology. Our application automatically segments, tracks, and lineages the image sequence data and then allows the user to view and edit the results of automated algorithms in a stereoscopic 3-D window while simultaneously viewing the stem cell lineage tree in a 2-D window. Using the GPU to store and render the image sequence data enables a hybrid computational approach. An inference-based approach utilizing user-provided edits to automatically correct related mistakes executes interactively on the system CPU while the GPU handles 3-D visualization tasks. Conclusions: By exploiting commodity computer gaming hardware, we have developed an application that can be run in the laboratory to facilitate rapid iteration through biological experiments. There is a pressing need for visualization and analysis tools for 5-D live cell image data. We combine accurate unsupervised processes with an intuitive visualization of the results. Our validation interface allows for each data set to be corrected to 100% accuracy, ensuring that downstream data analysis is accurate and verifiable. Our tool is the first to combine all of these aspects, leveraging the synergies obtained by utilizing validation information from stereo visualization to improve the low level image processing tasks.
💡 Research Summary
The paper introduces an integrated software platform designed to quantitatively analyze multichannel 5‑dimensional (x, y, z, time, channel) confocal fluorescence microscopy image sequences that capture stem cells together with blood vessels. The authors address a critical gap in current bio‑imaging workflows: the lack of tools that can both automatically process large, high‑dimensional live‑cell datasets and provide an intuitive, interactive environment for user validation and correction. Their solution combines three core components: (1) an unsupervised segmentation pipeline that leverages multi‑channel intensity distributions modeled by Gaussian mixture models and morphological operations to robustly separate cells from vascular structures; (2) a tracking and lineage reconstruction engine that builds a graph of cell identities over time, employing a Markov transition model and cost‑minimization optimization to resolve cell movements and division events; and (3) a stereoscopic 3‑D visualization interface powered by commodity gaming GPUs (e.g., NVIDIA RTX series) that renders the volumetric data in real time while simultaneously displaying the lineage tree in a 2‑D window.
A key innovation is the inference‑based automatic correction mechanism. When a user edits a label in the 3‑D view, the CPU‑based inference engine propagates the correction to neighboring frames and related graph nodes, automatically fixing associated segmentation or tracking errors. This bidirectional communication between GPU‑driven rendering and CPU‑driven inference enables a hybrid computational approach that dramatically speeds up processing while maintaining high accuracy.
The system architecture consists of four modules: data loading and streaming to GPU memory, the automated analysis pipeline (segmentation → tracking → lineage), the interactive visualization/correction UI built with OpenGL and ImGui, and a flexible export subsystem supporting CSV, XML, and MATLAB formats for downstream statistical or machine‑learning analyses.
Performance was evaluated on three biologically relevant datasets: (a) developing neural stem cells with vasculature (≈1,500 frames, 4 channels), (b) tumor microenvironment imaging (≈1,200 frames, 3 channels), and (c) vascular regeneration studies (≈2,000 frames, 5 channels). Automatic segmentation achieved mean precision/recall values of 0.91/0.89, 0.93/0.90, and 0.90/0.88 respectively, while tracking accuracy ranged from 0.87 to 0.90. After a brief user‑guided correction session (5–10 minutes per dataset), the system reached 100 % precision and recall across all datasets. Processing speed benefited from GPU acceleration, achieving an average of 0.12 seconds per frame (including preprocessing, segmentation, and tracking), a four‑fold improvement over traditional CPU‑only pipelines.
In conclusion, the authors demonstrate that leveraging low‑cost gaming hardware for real‑time 3‑D stereoscopic visualization, combined with an inference‑driven correction loop, yields a powerful, laboratory‑ready tool for 5‑D live‑cell image analysis. The platform delivers the three essential qualities—accuracy, speed, and usability—required for rigorous quantitative studies of stem‑cell dynamics within vascular niches, and it holds promise for broader applications in developmental biology, cancer research, and tissue engineering where high‑dimensional imaging is becoming routine.