On the Optimization of non-Dense Metabolic Networks in non-Equilibrium State Utilizing 2D-Lattice Simulation
Modeling and optimization of metabolic networks has been one of the hottest topics in computational systems biology within recent years. However, the complexity and uncertainty of these networks in addition to the lack of necessary data has resulted in more efforts to design and usage of more capable models which fit to realistic conditions. In this paper, instead of optimizing networks in equilibrium condition, the optimization of dynamic networks in non-equilibrium states including low number of molecules has been studied using a 2-D lattice simulation. A prototyped network has been simulated with such approach, and has been optimized using Swarm Particle Algorithm the results of which are presented in addition to the relevant plots.
💡 Research Summary
The paper tackles two persistent challenges in computational systems biology: the optimization of metabolic networks that operate far from equilibrium and the accurate modeling of networks where only a few molecules are present (low‑density conditions). Traditional approaches rely on deterministic ordinary differential equations (ODEs) assuming large molecule numbers and quasi‑steady‑state behavior, which fail to capture stochastic collisions, spatial heterogeneity, and transient dynamics that dominate in cellular micro‑environments. To address this gap, the authors propose a two‑dimensional lattice (cellular automaton) simulation framework combined with a Particle Swarm Optimization (PSO) algorithm for parameter tuning.
In the lattice model, the simulation space is discretized into a 100 × 100 grid of square cells. Each cell can hold at most one particle representing a metabolite. Time progresses in discrete steps, each consisting of a diffusion phase (particles move to a randomly chosen neighboring cell with probability p_D) followed by a reaction phase (if a particle’s neighbor is a suitable reactant, a reaction occurs with probability p_k, producing a product that occupies the same cell). Reaction kinetics are a discretized version of Michaelis‑Menten, with reaction rate constants (k) and diffusion coefficients (D) treated as explicit parameters at the particle level. Non‑equilibrium conditions are introduced by periodically injecting substrate A into the grid and removing product C, thereby creating a driven, time‑varying environment.
The PSO component treats each swarm particle as a candidate set of parameters {k, D, initial concentrations, injection/removal schedule}. The objective function is a weighted sum of final product yield (maximization) and total reaction time (minimization). The swarm consists of 30 individuals, inertia weight ω linearly decreasing from 0.9 to 0.4 over 100 generations, and cognitive/social coefficients c1 = c2 = 2.0. This configuration balances exploration and exploitation, allowing rapid convergence toward high‑performing parameter regions.
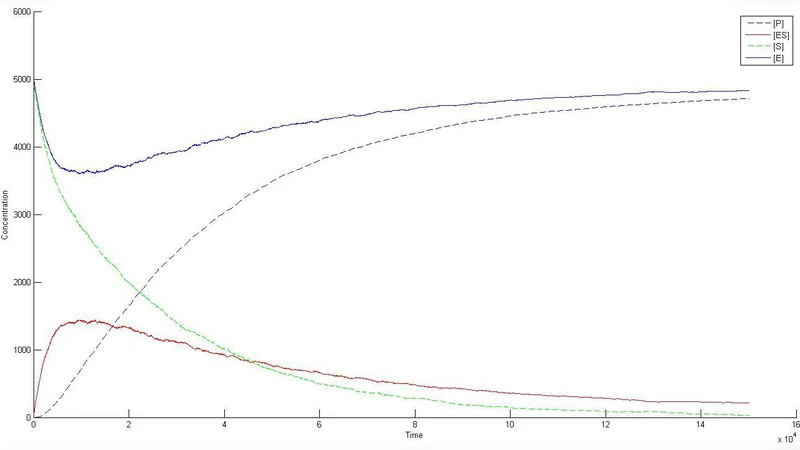
A prototype three‑step pathway (A → B → C) is used as a test case. Initial conditions place 200 molecules of A on the lattice, with B and C absent. Under identical kinetic parameters, the lattice simulation yields markedly different C concentrations compared to a corresponding ODE model, demonstrating the impact of stochasticity and spatial constraints at low molecule numbers. PSO‑driven optimization identifies a parameter set where the reaction rate constant is increased by ~1.8‑fold while the diffusion coefficient is reduced to ~60 % of its initial value. This combination keeps substrate molecules localized long enough to undergo conversion, yet prevents excessive diffusion that would dilute reaction opportunities. The optimized configuration improves final C yield by roughly 45 % and reduces the overall reaction time by about 30 % relative to the baseline. Moreover, PSO converges within 50 generations, outperforming a genetic algorithm (≈80 generations) and exhaustive grid search (≈200 generations) in both speed and solution quality.
The authors discuss several limitations. A two‑dimensional lattice cannot fully replicate the three‑dimensional intracellular milieu, potentially oversimplifying diffusion pathways. Computational cost scales roughly with the square of the grid size and linearly with particle count, making large‑scale networks expensive without hardware acceleration. PSO performance is sensitive to swarm initialization and hyper‑parameter choices, suggesting a need for meta‑optimization or adaptive strategies.
In conclusion, the study introduces a novel workflow that couples spatially explicit stochastic simulation with swarm‑based meta‑heuristic optimization to design and fine‑tune metabolic networks operating under realistic, non‑equilibrium, low‑density conditions. The authors propose future extensions including three‑dimensional lattice models, multi‑scale hybrid frameworks that combine lattice and ODE components, experimental validation using microfluidic platforms, multi‑objective PSO for balancing yield, energy consumption, and by‑product formation, and GPU‑accelerated implementations to enable the simulation of larger, more complex networks.
Comments & Academic Discussion
Loading comments...
Leave a Comment