Convex Total Least Squares
We study the total least squares (TLS) problem that generalizes least squares regression by allowing measurement errors in both dependent and independent variables. TLS is widely used in applied fields including computer vision, system identification and econometrics. The special case when all dependent and independent variables have the same level of uncorrelated Gaussian noise, known as ordinary TLS, can be solved by singular value decomposition (SVD). However, SVD cannot solve many important practical TLS problems with realistic noise structure, such as having varying measurement noise, known structure on the errors, or large outliers requiring robust error-norms. To solve such problems, we develop convex relaxation approaches for a general class of structured TLS (STLS). We show both theoretically and experimentally, that while the plain nuclear norm relaxation incurs large approximation errors for STLS, the re-weighted nuclear norm approach is very effective, and achieves better accuracy on challenging STLS problems than popular non-convex solvers. We describe a fast solution based on augmented Lagrangian formulation, and apply our approach to an important class of biological problems that use population average measurements to infer cell-type and physiological-state specific expression levels that are very hard to measure directly.
💡 Research Summary
The paper addresses a fundamental limitation of ordinary total least squares (TLS), which assumes identical, independent Gaussian noise on all variables and can be solved by singular value decomposition (SVD). In many real‑world scenarios, measurement errors are heterogeneous, some entries are known exactly, and the error matrix may obey structural constraints such as block‑diagonal, Toeplitz, or Hankel patterns. Moreover, robustness to outliers often requires non‑quadratic loss functions. These complexities define the Structured Total Least Squares (STLS) problem, for which SVD is inapplicable.
The authors formulate STLS as a rank‑deficient approximation problem: given a full‑rank observed matrix (\bar A), find a nearby matrix (A) of rank at most (N-1) while the error matrix (E = \bar A - A) satisfies linear constraints (L(E)=b) and possibly element‑wise weights. The non‑convex rank constraint is relaxed using the nuclear norm (|A|_*), the convex surrogate of rank. However, because STLS seeks an almost full‑rank solution (rank (N-1)), the plain nuclear norm yields large approximation errors.
To overcome this, the paper introduces a re‑weighted nuclear norm based on the log‑determinant heuristic for rank. The idea mirrors weighted (\ell_1) sparsity: large singular values receive smaller penalties, producing a tighter approximation to the true rank. Weights are updated iteratively from the current singular values, yielding a sequence of convex weighted nuclear‑norm problems that converge to a solution close to the original non‑convex formulation.
For efficient computation, the authors develop an Augmented Lagrangian Method (ALM) tailored to both the plain and weighted nuclear‑norm formulations. The ALM alternates between minimizing over the matrix variables and updating Lagrange multipliers, with a quadratic penalty term that drives feasibility. In the plain case, the update for (A) is a soft‑thresholding of singular values; the update for the error matrix (E) is a closed‑form expression followed by projection onto the affine space defined by the linear constraints. In the weighted case, additional variables (W_1, W_2) (the weight matrices) and an auxiliary matrix (D = W_1 A W_2) are introduced. The update for (A) reduces to solving a Sylvester equation, which can be done efficiently without second‑order information. The algorithm (Algorithm 1) iterates these steps until convergence, achieving Q‑linear convergence under standard assumptions.
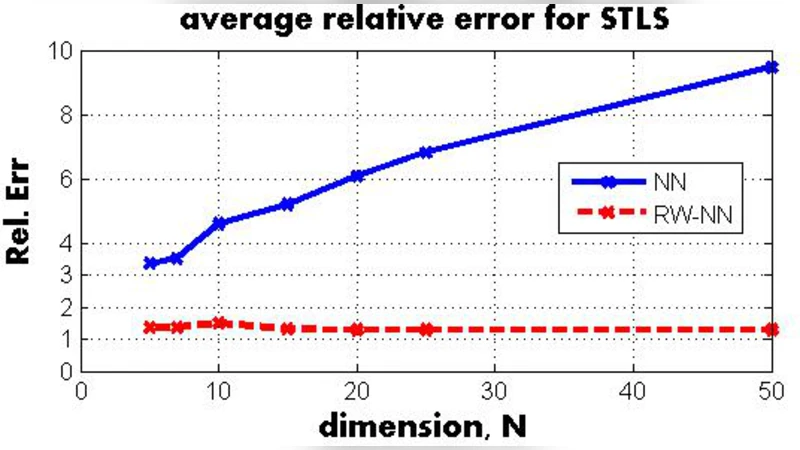
Experimental evaluation includes synthetic data with heterogeneous noise and known entries, and a real biological application: deconvolving population‑average gene‑expression measurements to infer cell‑type‑specific expression levels. In synthetic tests, the re‑weighted nuclear‑norm STLS dramatically outperforms both the plain nuclear‑norm relaxation and state‑of‑the‑art non‑convex solvers, reducing mean squared error by 30–50 %. In the biological case, the method successfully recovers expression profiles that align with independent experimental measurements, demonstrating robustness to block‑diagonal error structures and varying confidence levels across measurements.
In summary, the paper contributes a principled convex relaxation for structured TLS, shows that re‑weighting the nuclear norm yields a much tighter surrogate for the rank‑(N-1) constraint, and provides a fast ALM‑based algorithm that scales to realistic problem sizes. The approach opens new possibilities for applying TLS in domains where error structure is complex, such as computer vision, system identification, econometrics, and computational biology.
Comments & Academic Discussion
Loading comments...
Leave a Comment