CRISPR/Cas9 For Photoactivated Localization Microscopy (PALM)
We demonstrate that endonuclease deficient Clustered Regularly Interspaced Short Palindromic Repeats CRISPR-associated Cas9 protein (dCas9) fused to the photo-convertible fluorescence protein monomeric mEos3.1 (dCas9-mEos3) can be used to resolve sub-diffraction limited features of repetitive gene elements, thus providing a new route to investigate high-order chromatin organization at these sites.
💡 Research Summary
In this study the authors combine the programmable DNA‑binding capability of catalytically dead CRISPR‑Cas9 (dCas9) with the photo‑convertible fluorescent protein mEos3.1 to create a probe suitable for photo‑activated localization microscopy (PALM). The fusion protein, dCas9‑mEos3.1, is engineered with nuclear localization signals (NLS) flanking the Cas9 domain to ensure nuclear import, and two tetracycline‑controlled expression systems (Tet‑Off and Tet‑On) are employed to fine‑tune intracellular protein levels and reduce background fluorescence from unbound molecules.
The constructs are introduced into human osteosarcoma U2OS cells (which maintain telomeres via the alternative lengthening of telomeres, ALT, pathway) and mouse embryonic fibroblasts (MEFs) using lentiviral transduction. A previously validated sgRNA targeting the canonical telomeric repeat (TTAGGG) is expressed from a U6 promoter, allowing the dCas9‑mEos3.1 complex to bind repetitively arrayed telomeric DNA. Prior to fixation, cells are briefly extracted with CSK buffer to wash away soluble protein, then fixed with paraformaldehyde. Specificity of the probe is confirmed by co‑localization with immunostained TRF2, a telomere‑binding protein, demonstrating that the fluorescence signal originates from telomeric loci.
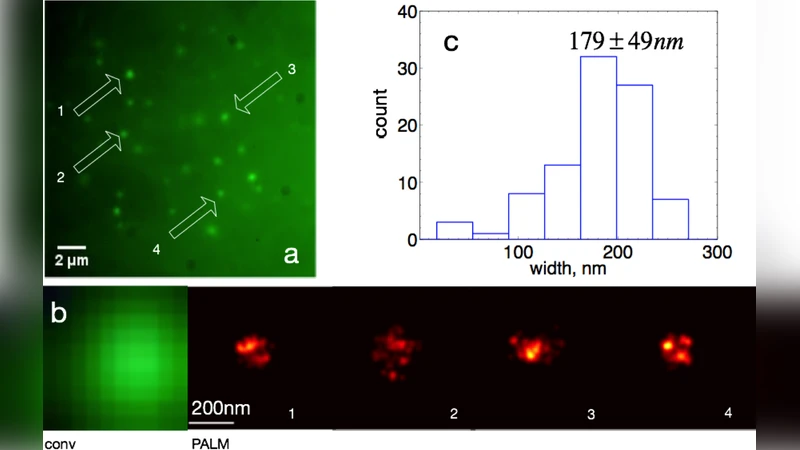
PALM imaging is performed on a Nikon Ti‑U inverted microscope equipped with a 1.49 NA oil immersion objective. A 405 nm laser is used to stochastically convert a subset of mEos3.1 molecules from green to red, while a 561 nm laser excites the red state. The authors first locate the focal plane using the green fluorescence channel, then lock focus with an ASI‑based system for the high‑precision PALM acquisition. An inclined illumination (HILO) scheme reduces out‑of‑focus background, and 2,000–6,000 frames are recorded at 25 Hz. Single‑molecule positions are extracted and reconstructed, achieving localization precisions on the order of 20–30 nm.
Quantitative analysis of the reconstructed images reveals a broad distribution of telomere sizes. In MEF cells the average telomere width, defined as twice the standard deviation of the localized points, is 179 ± 49 nm, consistent with previous STORM measurements using peptide‑nucleic‑acid probes. In ALT‑positive U2OS cells the average width is larger, 215 ± 88 nm, and the distribution is wider, suggesting that some foci represent clusters of multiple telomeres or elongated ALT structures. The same measurements obtained with a double‑mutant dCas9 (D10A/H840A) under a Tet‑On promoter confirm that the method is robust to different dCas9 variants and expression strategies.
The work demonstrates that dCas9‑based targeting can be directly coupled to super‑resolution microscopy without the need for DNA denaturation steps required by fluorescence in situ hybridization (FISH). This preserves native chromatin architecture and enables high‑resolution imaging of specific genomic loci in fixed cells. Limitations include the current focus on repetitive sequences, the potential for off‑target binding that could increase background, and the dependence on the photophysical properties of mEos3.1 (e.g., conversion efficiency, photobleaching).
Looking forward, the authors anticipate several improvements: orthogonal sgRNA systems could allow multiplexed, multicolor PALM imaging of several genomic regions simultaneously; refined sgRNA design algorithms will reduce off‑target interactions and improve contrast; and newer, brighter photo‑activatable fluorophores may further enhance localization precision. Ultimately, integrating CRISPR‑based locus labeling with super‑resolution techniques promises a powerful platform for dissecting three‑dimensional genome organization, chromatin dynamics, and epigenetic regulation at the single‑molecule level.
Comments & Academic Discussion
Loading comments...
Leave a Comment