A chromatin structure based model accurately predicts DNA replication timing in human cells
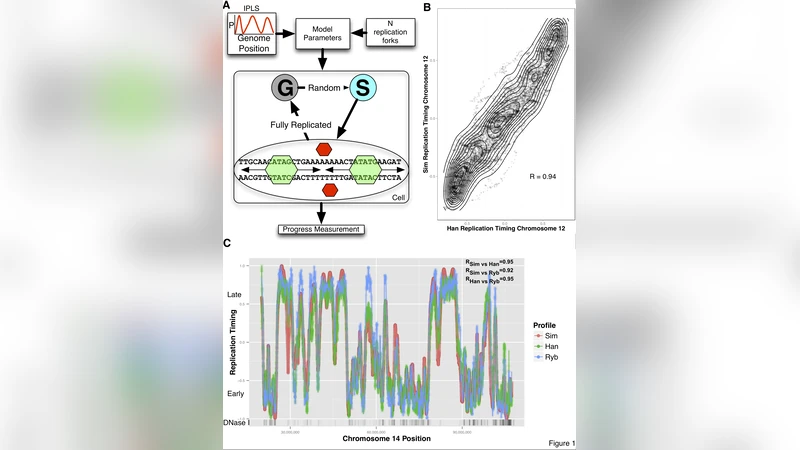
The metazoan genome is replicated in precise cell lineage specific temporal order. However, the mechanism controlling this orchestrated process is poorly understood as no molecular mechanisms have been identified that actively regulate the firing sequence of genome replication. Here we develop a mechanistic model of genome replication capable of predicting, with accuracy rivaling experimental repeats, observed empirical replication timing program in humans. In our model, replication is initiated in an uncoordinated (time-stochastic) manner at well-defined sites. The model contains, in addition to the choice of the genomic landmark that localizes initiation, only a single adjustable parameter of direct biological relevance: the number of replication forks. We find that DNase hypersensitive sites are optimal and independent determinants of DNA replication initiation. We demonstrate that the DNA replication timing program in human cells is a robust emergent phenomenon that, by its very nature, does not require a regulatory mechanism determining a proper replication initiation firing sequence.
💡 Research Summary
The authors present a parsimonious mechanistic model that accurately predicts genome‑wide DNA replication timing in human cells. The model treats replication initiation as a stochastic, time‑independent process in which a limited number of replication forks (the sole adjustable biological parameter) independently select unreplicated genomic loci according to an Initiation Probability Landscape (IPLS). To identify the IPLS that best recapitulates observed timing profiles, the authors evaluated 167 ENCODE-derived genomic marks across multiple cell lines. DNase hypersensitive (HS) sites emerged as the most powerful predictor (Pearson r = 0.87), outperforming static features such as transcription‑start sites, CpG islands, GC content, and even other dynamic chromatin marks. When co‑localized marks were removed, only DNase HS retained predictive power, indicating it is the primary independent determinant of replication timing.
The model’s single biological parameter—the maximum number of simultaneously active replication forks (N)—was optimized per chromosome and found to scale linearly with chromosome length (≈1 fork per 1.3 Mb). Using this scaling, simulated S‑phase duration (≈8.3 h) matches experimentally measured values (6–10 h). Incorporating a technical flow‑sorting variable further improved correlation with experimental data to r = 0.92, approaching the inter‑experimental reproducibility limit (r = 0.94).
The framework was validated across cell types (fibroblasts, lymphoblastoid, leukemia), species (human and mouse fibroblasts), and genomic rearrangements (in silico t(12;21) translocation), consistently reproducing lineage‑specific timing patterns and the abrupt timing shifts observed near chromosomal breakpoints. Robustness analyses showed that removing up to 75 % of DNase HS sites or altering the functional mapping of ENCODE signal amplitudes had minimal impact on predictive accuracy, underscoring that the spatial distribution of DNase HS sites—not the precise initiation probabilities—drives the emergent replication timing program.
Overall, the study demonstrates that DNA replication timing in metazoan genomes is an emergent, systems‑level property dictated chiefly by chromatin accessibility (as marked by DNase hypersensitivity) and the global limit on replication fork number, without requiring a dedicated regulatory mechanism to order origin firing. This minimal model provides a powerful tool for predicting replication dynamics in diverse cellular contexts and offers new insights into how chromatin architecture shapes genome replication.
Comments & Academic Discussion
Loading comments...
Leave a Comment