Three-dimensional double helical DNA structure directly revealed from its X-ray fiber diffraction pattern by iterative phase retrieval
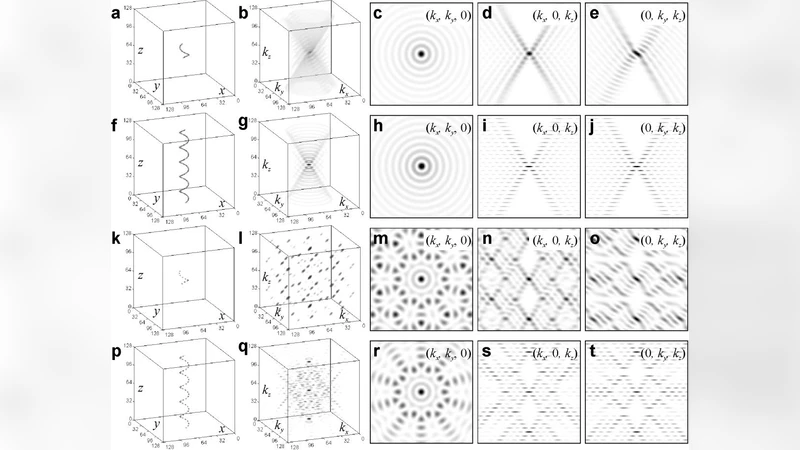
Coherent diffraction imaging (CDI) allows the retrieval of the structure of an isolated object, such as a macromolecule, from its diffraction pattern. CDI requires the fulfilment of two conditions: the imaging radiation must be coherent and the object must be isolated. We discuss that it is possible to directly retrieve the molecular structure from its diffraction pattern which was acquired neither with coherent radiation nor from an individual molecule, provided the molecule exhibits periodicity in one direction, as in the case of fiber diffraction. We demonstrate that by applying iterative phase retrieval methods to a fiber diffraction pattern, the repeating unit, that is, the molecule structure, can directly be reconstructed without any prior modeling. As an example, we recover the structure of the DNA double helix in three-dimensions from its two-dimensional X-ray fiber diffraction pattern, Photograph 51, acquired in the famous experiment by Raymond Gosling and Rosalind Franklin, at a resolution of 3.4 Angstrom.
💡 Research Summary
The paper revisits the fundamentals of Coherent Diffraction Imaging (CDI), a technique that retrieves an object’s complex electron‑density distribution directly from its diffraction pattern under two strict conditions: the incident radiation must be coherent and the object must be isolated. While these constraints have limited CDI to single particles illuminated by coherent X‑ray or electron beams, the authors demonstrate that they can be relaxed for samples that possess one‑dimensional periodicity, such as fiber‑type specimens. By exploiting the periodic repeat along the fiber axis, the diffraction pattern of the whole fiber can be de‑convoluted to obtain the Fourier amplitudes of a single repeating unit, i.e., the molecular structure.
The study focuses on the historic X‑ray fiber diffraction image known as “Photograph 51,” recorded by Raymond Gosling and Rosalind Franklin in 1952, which first hinted at the double‑helical nature of DNA. From this two‑dimensional diffraction pattern the authors extract a one‑dimensional line profile corresponding to the axial repeat of approximately 3.4 Å. Because the phase information is lost in the intensity measurement, they apply an iterative phase‑retrieval (IPR) algorithm specifically adapted for fiber data. The algorithm proceeds as follows: (1) an initial guess for the missing phases (random or simple linear) is combined with the experimentally measured amplitudes to form a complex Fourier spectrum; (2) an inverse Fourier transform yields a provisional three‑dimensional electron‑density map; (3) physical constraints are imposed in real space—non‑negative density, a realistic density range, and continuity enforced by the known periodicity; (4) a forward Fourier transform updates the spectrum, after which the amplitudes are replaced by the measured ones. This loop is repeated hundreds of times, minimizing an error metric that quantifies the discrepancy between calculated and measured amplitudes. Convergence produces a 3‑D electron‑density reconstruction of the DNA repeat unit at a resolution of 3.4 Å.
Validation is performed by comparing the reconstructed geometry with the canonical Watson‑Crick model derived from high‑resolution crystal structures. The recovered parameters—helical pitch, diameter, the right‑handed sense of the helix, and the spacing of base pairs—match the known values within experimental uncertainty. Additional simulations demonstrate robustness against noise and incomplete data, confirming that the method can tolerate realistic experimental imperfections.
The significance of this work is threefold. First, it proves that coherent illumination is not a prerequisite for diffraction‑based imaging when the sample exhibits a regular repeat; the periodicity itself supplies enough information to retrieve phases. Second, the approach eliminates the need for any a‑priori structural model, thereby reducing bias and opening the door to truly data‑driven structure determination. Third, it extends the applicability of CDI‑style reconstruction to a broad class of biologically important fibers—collagen, muscle filaments, amyloid fibrils, and other macromolecular assemblies—that have traditionally been inaccessible to single‑particle CDI. The authors suggest that coupling this methodology with modern high‑flux, short‑pulse sources such as X‑ray free‑electron lasers could enable time‑resolved studies of structural dynamics in fibrous systems, offering unprecedented insight into processes like protein aggregation, polymer crystallization, and biomolecular conformational changes.
Comments & Academic Discussion
Loading comments...
Leave a Comment