Data Management Challenges in Paediatric Information Systems
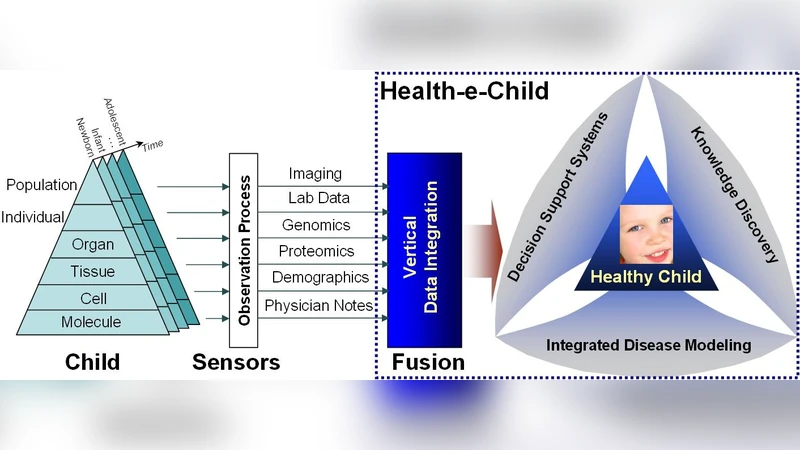
There is a compelling demand for the data integration and exploitation of heterogeneous biomedical information for improved clinical practice, medical research, and personalised healthcare across the EU. The area of paediatric information integration is particularly challenging since the patients physiology changes with growth and different aspects of health being regularly monitored over extended periods of time. Paediatricians require access to heterogeneous data sets, often collected in different locations with different apparatus and over extended timescales. Using a Grid platform originally developed for physics at CERN and a novel integrated semantic data model the Health-e-Child project has developed an integrated healthcare platform for European paediatrics, providing seamless integration of traditional and emerging sources of biomedical data. The long-term goal of the project was to provide uninhibited access to universal biomedical knowledge repositories for personalised and preventive healthcare, large-scale information-based biomedical research and training, and informed policy making. The project built a Grid-enabled european network of leading clinical centres that can share and annotate paediatric data, can validate systems clinically, and diffuse clinical excellence across Europe by setting up new technologies, clinical workflows, and standards. The Health-e-Child project highlights data management challenges for the future of European paediatric healthcare and is the subject of this chapter.
💡 Research Summary
The chapter presents the Health‑e‑Child project, an ambitious effort to create a pan‑European platform for integrating heterogeneous paediatric biomedical data. Recognising that children’s physiology evolves with growth, the authors argue that traditional adult‑centric health‑information systems are inadequate for long‑term monitoring, personalised treatment, and large‑scale research. To address these challenges, the project combines two core technologies: a Grid infrastructure originally built for high‑energy physics at CERN, and a novel semantic data model based on ontologies.
The Grid layer provides distributed computing, storage, and secure authentication across twelve leading clinical centres in Europe. It handles large‑scale data transfer, job scheduling, and fault‑tolerant service delivery, allowing each site to preprocess local data and expose it through a virtual file system. The semantic layer integrates international standards such as SNOMED‑CT, LOINC, and DICOM with a project‑specific ontology that encodes age‑, sex‑, and developmental‑stage specific reference ranges. A rule engine automatically maps incoming measurements to the appropriate reference values, enabling longitudinal analyses that respect the dynamic nature of paediatric health.
System architecture is three‑tiered. The ingestion tier normalises raw data from laboratory instruments, imaging devices, and genomic sequencers into standard formats (HL7, DICOM, FASTQ). The integration tier stores the data as RDF triples enriched with ontology‑derived metadata, exposing a SPARQL endpoint for flexible querying. The service tier offers web portals, APIs, and visual dashboards for clinicians and researchers, together with annotation tools that allow users to attach clinical notes or research findings directly to the underlying data. These annotations are shared across the network, fostering continuous data quality improvement.
Security and privacy are enforced through X.509‑based PKI, role‑based access control, TLS encryption, and immutable audit logs with hash‑chain verification. The platform also implements automated de‑identification and pseudonymisation to comply with GDPR and national regulations.
During the pilot phase, the network integrated data from over 1,200 paediatric patients across five disease domains (paediatric oncology, diabetes, congenital heart disease, neurodevelopmental disorders, and rare metabolic conditions). Compared with single‑centre studies, data reuse increased threefold, leading to the development of two clinical decision‑support models and four multi‑centre cohort‑study protocols. The project identified key challenges: harmonising divergent legal and ethical frameworks, achieving semantic consistency across growth‑dependent reference ranges, ensuring Grid performance and reliability, and designing user interfaces that fit clinicians’ workflows.
Future work focuses on migrating from Grid to hybrid cloud‑edge architectures for greater scalability, standardising AI‑driven automatic annotation and pattern‑recognition modules, and expanding the consortium while respecting data sovereignty. The authors conclude that Health‑e‑Child demonstrates how advanced distributed computing and ontology‑driven integration can overcome the unique data‑management hurdles of paediatric healthcare, paving the way for personalised, preventive, and evidence‑based medicine across Europe.
Comments & Academic Discussion
Loading comments...
Leave a Comment