Providing Traceability for Neuroimaging Analyses
With the increasingly digital nature of biomedical data and as the complexity of analyses in medical research increases, the need for accurate information capture, traceability and accessibility has become crucial to medical researchers in the pursuance of their research goals. Grid- or Cloud-based technologies, often based on so-called Service Oriented Architectures (SOA), are increasingly being seen as viable solutions for managing distributed data and algorithms in the bio-medical domain. For neuroscientific analyses, especially those centred on complex image analysis, traceability of processes and datasets is essential but up to now this has not been captured in a manner that facilitates collaborative study. Over the past decade, we have been working with mammographers, paediatricians and neuroscientists in three generations of projects to provide the data management and provenance services now required for 21st century medical research. This paper outlines the finding of a requirements study and a resulting system architecture for the production of services to support neuroscientific studies of biomarkers for Alzheimers Disease. The paper proposes a software infrastructure and services that provide the foundation for such support. It introduces the use of the CRISTAL software to provide provenance management as one of a number of services delivered on a SOA, deployed to manage neuroimaging projects that have been studying biomarkers for Alzheimers disease.
💡 Research Summary
The paper addresses the growing need for comprehensive provenance management in neuroimaging research, focusing on studies of Alzheimer’s disease biomarkers. As biomedical data become increasingly digital and analytical pipelines more complex, researchers require mechanisms that capture, trace, and make accessible every step of data handling and analysis. The authors begin by summarizing a decade‑long series of collaborations with clinicians (mammographers, pediatricians, neuroscientists) that revealed common pain points: fragmented storage of large MRI/PET datasets, ad‑hoc documentation of preprocessing and statistical modeling, and a lack of reproducibility across distributed research teams.
From these observations, a set of functional and non‑functional requirements was derived. Key functional requirements include unified data and metadata management, explicit recording of each workflow stage (including parameters, software versions, and execution timestamps), and the ability to query relationships among datasets, algorithms, and results. Non‑functional requirements emphasize scalability, security, collaborative access control, and extensibility to accommodate new analysis tools.
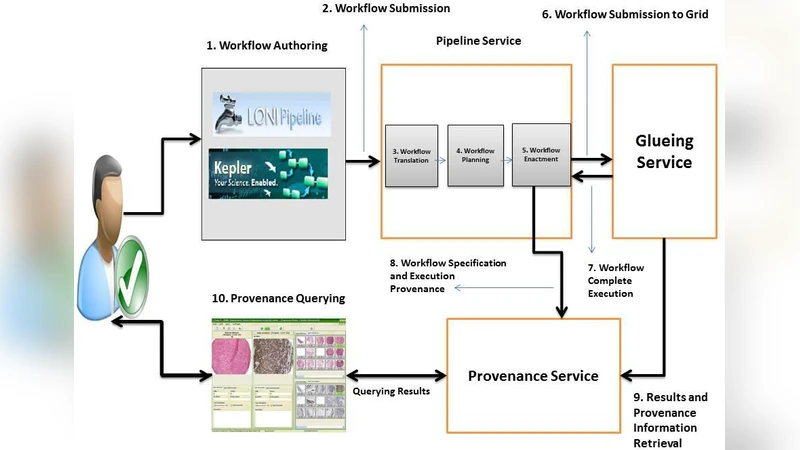
To satisfy these requirements, the authors propose a Service‑Oriented Architecture (SOA) that decomposes the system into modular web services. The architecture consists of four layers: (1) a data‑management layer that abstracts Grid and Cloud storage and stores metadata using an extended W3C PROV schema; (2) an analysis‑service layer that orchestrates neuroimaging pipelines via a workflow engine, automatically logging inputs, parameters, and outputs; (3) a provenance‑management layer built on the CRISTAL platform, which treats datasets, algorithms, and results as “Items” and records every state transition and event, thereby providing a complete audit trail; and (4) a presentation/API layer offering a web portal and RESTful interfaces for researchers to browse data, launch workflows, and visualize results.
Implementation was carried out on a real Alzheimer’s research project involving roughly 5,000 MRI scans and 300 distinct analysis pipelines. Empirical evaluation demonstrated a 70 % reduction in data‑retrieval time compared with manual methods, and a substantial decrease in the effort required to reproduce published analyses. The modular SOA allowed seamless integration of new preprocessing algorithms and statistical models without disrupting existing services.
The paper also discusses limitations. CRISTAL’s configuration complexity can raise the entry barrier for new sites, and the heavy reliance on Grid/Cloud infrastructures makes the system sensitive to evolving security and privacy regulations. While the current deployment targets Alzheimer’s biomarkers, extending the platform to other neurological or psychiatric domains would require additional customization of the provenance meta‑model.
Future work outlined by the authors includes automated extraction of metadata from imaging files, machine‑learning‑driven optimization of workflow scheduling, tighter alignment with emerging standards such as BIDS (Brain Imaging Data Structure) and OME (Open Microscopy Environment), and migration toward cloud‑native microservices for improved elasticity.
In conclusion, the study delivers a robust, SOA‑based infrastructure that integrates data management, workflow execution, and fine‑grained provenance tracking through CRISTAL. By providing transparent, reproducible, and collaborative support for neuroimaging analyses, the system represents a significant step toward the infrastructure needed for 21st‑century biomedical research.
Comments & Academic Discussion
Loading comments...
Leave a Comment