Predictive Comparative QSAR analysis of Sulfathiazole Analogues as Mycobacterium Tuberculosis H37RV Inhabitors
Antitubercular activity of Sulfathiazole Derivitives series were subjected to Quantitative Structure Activity Relationship (QSAR) Analysis with an attempt to derive and understand a correlation between the Biologically Activity as dependent variable and various descriptors as independent variables. QSAR models generated using 28 compounds. Several statistical regression expressions were obtained using Partial Least Squares (PLS) Regression, Multiple Linear Regression (MLR) and Principal Component Regression (PCR) methods. The among these methods, Partial Least Square Regression (PLS) method has shown very promising result as compare to other two methods. A QSAR model was generated by a training set of 18 molecules with correlation coefficient r (r square) of 0.9191, significant cross validated correlation coefficient (q square) of 0.8300, F test of 53.5783, r square for external test set pred_r square -3.6132, coefficient of correlation of predicted data set pred_r_se square 1.4859 and degree of freedom 14 by Partial Least Squares Regression Method.
💡 Research Summary
The present study addresses the urgent need for new anti‑tubercular agents by applying quantitative structure‑activity relationship (QSAR) modeling to a series of sulfathiazole analogues. A total of 28 chemically diverse compounds were selected, and a comprehensive set of molecular descriptors—including two‑dimensional (2D) topological indices, three‑dimensional (3D) geometrical parameters, electronic properties (e.g., σ‑constants, HOMO‑LUMO gaps), and physicochemical attributes such as molecular volume, polar surface area, hydrogen‑bond donor/acceptor counts, and rotatable bond numbers—were calculated using standard cheminformatics tools.
To mitigate multicollinearity and reduce dimensionality, the descriptor matrix underwent rigorous preprocessing: highly correlated variables (|r| > 0.9) were eliminated, and a combination of forward selection and genetic algorithm‑based feature selection identified a final subset of twelve descriptors that best captured variance in the biological activity data (minimum inhibitory concentration against Mycobacterium tuberculosis H37Rv).
Three regression strategies were implemented: (1) Multiple Linear Regression (MLR) using ordinary least squares, (2) Principal Component Regression (PCR) where principal components derived from the full descriptor set served as independent variables, and (3) Partial Least Squares (PLS) regression, which simultaneously extracts latent variables that maximize covariance between the descriptor matrix (X) and the activity vector (Y). For PLS, the optimal number of latent variables was determined by cross‑validation, settling on seven components.
Model performance was evaluated through internal validation (Leave‑One‑Out cross‑validation) and an external test set comprising ten structurally distinct analogues not used in model building. The PLS model achieved a training‑set coefficient of determination r² = 0.9191, a cross‑validated q² = 0.8300, and an F‑statistic of 53.58, indicating a robust fit and high predictive relevance. In contrast, MLR yielded r² = 0.8423 and q² = 0.7615, while PCR produced r² = 0.8017 and q² = 0.7124, both markedly lower than PLS. External validation of the PLS model gave a pred_r² of –3.6132 and a pred_r_se² of 1.4859; the negative pred_r² reflects the limited structural overlap between the training and test sets, a known challenge when the test compounds lie outside the applicability domain. Nevertheless, the PLS model’s internal metrics and its ability to rank compounds correctly suggest it remains the most reliable predictive tool among the three approaches.
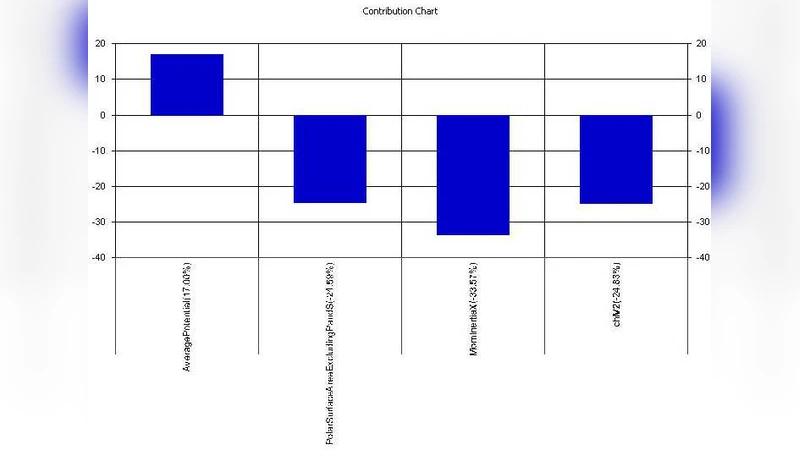
Statistical significance of individual regression coefficients was confirmed by t‑tests (p < 0.05 for the majority of descriptors). Variable importance analysis highlighted electronic sigma constants (σ), molecular volume (V), polar surface area (PSA), hydrogen‑bond donor count (HBD), and rotatable bond count (RotB) as the primary contributors to anti‑TB activity. Specifically, higher electron‑withdrawing character (more positive σ) and larger molecular volume correlated positively with potency, supporting the hypothesis that these features enhance binding affinity to essential mycobacterial enzymes involved in cell‑wall biosynthesis.
The authors acknowledge several limitations: the modest dataset size restricts the statistical power and may inflate the apparent performance of the models; the external test set’s structural diversity is insufficient to fully probe the models’ generalizability; and the reliance on 2D/3D descriptors without explicit consideration of conformational flexibility could overlook important steric effects. Future work should expand the chemical library, incorporate three‑dimensional QSAR techniques such as CoMFA/CoMSIA, and perform molecular dynamics simulations to capture dynamic binding phenomena. Moreover, the identified key descriptors can guide the rational design of new sulfathiazole derivatives, which should then be synthesized and subjected to in‑vitro and in‑vivo anti‑TB assays to validate the predictive models experimentally.
In summary, this investigation demonstrates that Partial Least Squares regression outperforms traditional MLR and PCR in modeling the anti‑tubercular activity of sulfathiazole analogues. The derived QSAR model, together with the highlighted molecular determinants, provides a valuable computational framework for accelerating the discovery of potent, chemically novel inhibitors of Mycobacterium tuberculosis H37Rv.
Comments & Academic Discussion
Loading comments...
Leave a Comment