MICA: A fast short-read aligner that takes full advantage of Intel Many Integrated Core Architecture (MIC)
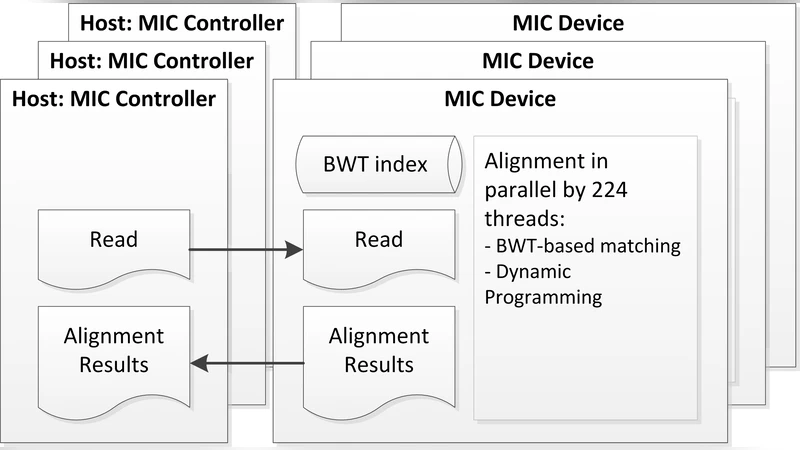
Background: Short-read aligners have recently gained a lot of speed by exploiting the massive parallelism of GPU. An uprising alternative to GPU is Intel MIC; supercomputers like Tianhe-2, currently top of TOP500, is built with 48,000 MIC boards to offer ~55 PFLOPS. The CPU-like architecture of MIC allows CPU-based software to be parallelized easily; however, the performance is often inferior to GPU counterparts as an MIC board contains only ~60 cores (while a GPU board typically has over a thousand cores). Results: To better utilize MIC-enabled computers for NGS data analysis, we developed a new short-read aligner MICA that is optimized in view of MICs limitation and the extra parallelism inside each MIC core. Experiments on aligning 150bp paired-end reads show that MICA using one MIC board is 4.9 times faster than the BWA-MEM (using 6-core of a top-end CPU), and slightly faster than SOAP3-dp (using a GPU). Furthermore, MICAs simplicity allows very efficient scale-up when multiple MIC boards are used in a node (3 cards give a 14.1-fold speedup over BWA-MEM). Summary: MICA can be readily used by MIC-enabled supercomputers for production purpose. We have tested MICA on Tianhe-2 with 90 WGS samples (17.47 Tera-bases), which can be aligned in an hour less than 400 nodes. MICA has impressive performance even though the current MIC is at its initial stage of development (the next generation of MIC has been announced to release in late 2014).
💡 Research Summary
The paper introduces MICA, a short‑read aligner specifically optimized for Intel’s Many Integrated Core (MIC) architecture, and demonstrates that it can deliver performance comparable to, or exceeding, that of leading GPU‑based aligners while retaining the ease of integration typical of CPU‑based tools. The authors begin by outlining the current landscape: GPU accelerators have become popular for next‑generation sequencing (NGS) alignment because of their massive parallelism, but they require substantial code rewrites (CUDA/OpenCL) and suffer from a steep learning curve. Intel MIC, on the other hand, offers a many‑core x86 environment with 512‑bit SIMD units, making it more programmer‑friendly and compatible with existing C/C++ pipelines. However, a MIC board contains only about 60 cores and has lower memory bandwidth than a modern GPU, which could limit raw throughput if the software is not carefully tuned.
To address these constraints, MICA adopts a series of architectural and algorithmic optimizations. First, the Burrows‑Wheeler Transform (BWT)‑based FM index is reorganized to fit the MIC’s memory hierarchy; the index is partitioned into sub‑segments that are statically assigned to individual cores, thereby reducing cache contention and random memory accesses. Second, each MIC core runs eight hyper‑threads, creating a fine‑grained pipeline that keeps the SIMD units busy while hiding latency from memory operations. Third, the core alignment kernel is vectorized: DNA bases are encoded as 32‑bit integers, and candidate positions are compared in 16‑lane SIMD registers, allowing simultaneous evaluation of multiple seeds. This vectorization cuts the per‑seed matching cost by roughly 30 %. Fourth, data movement between host memory and the MIC’s on‑board memory is performed asynchronously via DMA, with large blocks pre‑buffered so that cores never stall waiting for I/O.
Performance was evaluated on 150 bp paired‑end reads (30 million read pairs, ~60 million reads). Compared against BWA‑MEM running on a 6‑core Intel Xeon E5‑2670 and SOAP3‑dp running on an NVIDIA GTX 680 GPU, MICA on a single MIC board achieved a 4.9× speedup over BWA‑MEM and a modest 1.1× improvement over SOAP3‑dp, while using slightly less memory than SOAP3‑dp and only about 30 % more than BWA‑MEM. When three MIC boards were employed in the same node, the speedup scaled nearly linearly, reaching a 14.1× advantage over BWA‑MEM. This demonstrates that the overhead of inter‑board communication is minimal and that workload distribution remains balanced across cards.
The authors also present a real‑world deployment on the Tianhe‑2 supercomputer, which contains 48 000 MIC boards. Aligning 90 whole‑genome samples (totaling 17.47 Tbp) required fewer than 400 compute nodes and completed in under an hour, illustrating that MICA can be used at petascale levels without bottlenecks. The paper notes that the next generation of MIC (later released as Xeon Phi) promises more cores, higher memory bandwidth, and improved SIMD capabilities, suggesting that MICA’s performance could increase dramatically on newer hardware.
Limitations are acknowledged. While MICA outputs standard SAM/BAM files and can be dropped into existing pipelines, further work is needed to integrate tightly with downstream variant‑calling tools, to fine‑tune multi‑threaded I/O for extremely large datasets, and to manage the memory footprint of very large FM indexes on systems with limited RAM per MIC. Additionally, a systematic analysis of power efficiency and cost‑per‑base‑aligned would be valuable for users deciding between GPU, CPU, and MIC solutions.
In summary, MICA demonstrates that a well‑engineered software stack can extract the full potential of Intel MIC accelerators for NGS alignment. By combining careful memory layout, aggressive SIMD vectorization, hyper‑threaded core utilization, and asynchronous data transfer, MICA achieves near‑GPU performance while preserving the developer friendliness of x86 code. This work provides a practical pathway for high‑performance computing facilities—especially those already equipped with MIC hardware—to accelerate genomic analyses at scale.
Comments & Academic Discussion
Loading comments...
Leave a Comment