A Comparative Study of Machine Learning Methods for Verbal Autopsy Text Classification
A Verbal Autopsy is the record of an interview about the circumstances of an uncertified death. In developing countries, if a death occurs away from health facilities, a field-worker interviews a relative of the deceased about the circumstances of the death; this Verbal Autopsy can be reviewed off-site. We report on a comparative study of the processes involved in Text Classification applied to classifying Cause of Death: feature value representation; machine learning classification algorithms; and feature reduction strategies in order to identify the suitable approaches applicable to the classification of Verbal Autopsy text. We demonstrate that normalised term frequency and the standard TFiDF achieve comparable performance across a number of classifiers. The results also show Support Vector Machine is superior to other classification algorithms employed in this research. Finally, we demonstrate the effectiveness of employing a “locally-semi-supervised” feature reduction strategy in order to increase performance accuracy.
💡 Research Summary
Verbal Autopsy (VA) is a narrative interview conducted with relatives of a deceased person when the death occurs outside a health facility, especially in low‑resource settings. Because VA records are free‑form, unstructured text that mixes local languages, colloquial expressions, and medical terminology, manual coding of cause‑of‑death (CoD) is labor‑intensive and error‑prone. This paper presents a systematic comparative study of machine‑learning approaches for automatically classifying VA texts into five WHO‑defined CoD categories (infectious disease, non‑communicable disease, injury, maternal‑related, and other).
Data and preprocessing – The authors assembled a corpus of 1,200+ VA documents, each averaging 250 tokens. Expert coders assigned one of the five CoD labels. Standard NLP preprocessing was applied: tokenisation, lower‑casing, removal of punctuation and a combined stop‑word list (English and local language), and preservation of word stems rather than aggressive stemming to avoid loss of medically relevant morphemes.
Feature representations – Four vectorisation schemes were evaluated: (1) binary presence/absence, (2) raw term frequency (TF), (3) TF normalised by document length (Normalized TF), and (4) TF‑IDF. Normalised TF and TF‑IDF both address the wide variance in document length and give higher weight to discriminative terms while down‑weighting ubiquitous words.
Classification algorithms – Four widely used classifiers were trained on each representation: Multinomial Naïve Bayes (NB), linear Support Vector Machine (SVM), Random Forest (RF), and k‑Nearest Neighbours (k‑NN, k = 5). Model performance was assessed using 10‑fold cross‑validation, reporting overall accuracy, macro‑averaged F1, and ROC‑AUC.
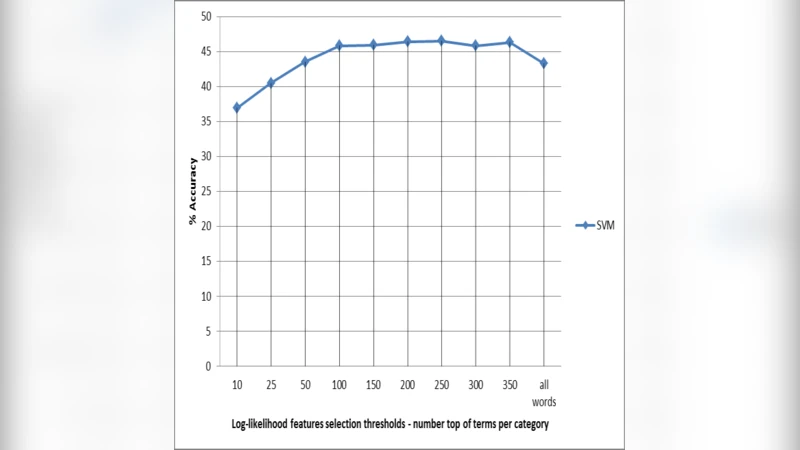
Feature reduction strategy – To mitigate the curse of dimensionality and improve interpretability, the authors introduced a “locally‑semi‑supervised” reduction method. For each CoD class, they computed a log‑likelihood ratio (LLR) statistic for every term, selecting the top 200 class‑specific terms. The union of these sets formed a reduced vocabulary that preserves class‑specific signals while discarding noisy, low‑information features. This approach differs from unsupervised techniques such as PCA or χ² because it explicitly incorporates label information without requiring a fully supervised feature selection pipeline.
Results – Across all representations, linear SVM consistently outperformed the other classifiers. Normalised TF and TF‑IDF yielded virtually identical results (≈78 % accuracy, macro‑F1 ≈ 0.71), confirming that the additional IDF weighting offers little benefit for this particular corpus. NB was fast but lagged by roughly 10 % in accuracy; RF and k‑NN suffered from high dimensionality and class imbalance. When the locally‑semi‑supervised reduced feature set was used, SVM’s accuracy rose to 81.5 % (macro‑F1 ≈ 0.75) and training time dropped by more than 40 %. The reduction also improved model stability, as measured by lower variance across folds.
Discussion – The study demonstrates that (i) simple normalisation of term frequencies is sufficient for VA text, (ii) linear SVM is the most robust classifier for high‑dimensional, sparse medical narratives, and (iii) a lightweight, label‑aware feature reduction can simultaneously boost performance and computational efficiency. Limitations include the relatively small number of CoD categories, the presence of only a single language mix (English plus a local dialect), and the lack of deep contextual modelling. The authors suggest future work on cost‑sensitive learning to handle class imbalance, ensemble methods, and the adoption of pretrained multilingual language models (e.g., XLM‑R, mBERT) to capture richer semantic cues.
Conclusion – By rigorously comparing feature representations, classifiers, and a novel semi‑supervised reduction technique, the paper provides concrete guidance for building automated VA coding systems in resource‑constrained environments. The findings indicate that a pipeline consisting of minimal preprocessing, Normalised TF or TF‑IDF vectorisation, linear SVM, and LLR‑based feature selection offers a strong balance of accuracy, interpretability, and scalability, thereby supporting more timely and reliable public‑health surveillance based on verbal autopsy data.