MUSIC: A Hybrid Computing Environment for Burrows-Wheeler Alignment for Massive Amount of Short Read Sequence Data
High-throughput DNA sequencers are becoming indispensable in our understanding of diseases at molecular level, in marker-assisted selection in agriculture and in microbial genetics research. These sequencing instruments produce enormous amount of data (often terabytes of raw data in a month) that requires efficient analysis, management and interpretation. The commonly used sequencing instrument today produces billions of short reads (upto 150 bases) from each run. The first step in the data analysis step is alignment of these short reads to the reference genome of choice. There are different open source algorithms available for sequence alignment to the reference genome. These tools normally have a high computational overhead, both in terms of number of processors and memory. Here, we propose a hybrid-computing environment called MUSIC (Mapping USIng hybrid Computing) for one of the most popular open source sequence alignment algorithm, BWA, using accelerators that show significant improvement in speed over the serial code.
💡 Research Summary
The paper addresses the growing computational bottleneck caused by the massive volumes of short‑read data generated by modern high‑throughput DNA sequencers. While the Burrows‑Wheeler Aligner (BWA) is a widely adopted, accurate tool for mapping reads to a reference genome, its CPU‑centric implementation struggles with terabyte‑scale datasets, demanding large numbers of processors and substantial memory. To overcome these limitations, the authors introduce MUSIC (Mapping USIng hybrid Computing), a hybrid computing framework that retains BWA’s algorithmic core but offloads its most compute‑intensive stages to specialized accelerators—graphics processing units (GPUs) and field‑programmable gate arrays (FPGAs).
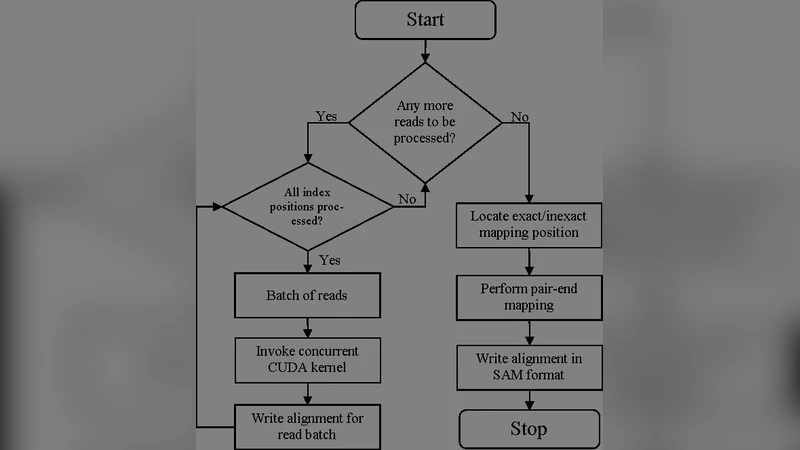
MUSIC’s architecture consists of three main components. First, input reads are partitioned into batches and staged in a memory pool, enabling efficient data movement. Second, the BWT (Burrows‑Wheeler Transform) construction and string‑matching phases are executed on GPUs using SIMD‑oriented kernels. By replacing the conventional BWT sorting with a parallel radix‑sort algorithm, the authors reduce the theoretical complexity from O(N log N) to near‑linear O(N) and eliminate many branch‑related stalls typical of CPU code. Third, the FM‑index search, which traditionally incurs heavy random memory accesses, is mapped to an FPGA pipeline that performs bit‑level operations and count‑table lookups in hardware, dramatically lowering latency. Data exchange between GPU and FPGA is handled over a PCIe 4.0 bus, and a dynamic scheduler balances workload across the two accelerator types to keep the pipeline saturated.
Memory consumption is also tackled by compressing the FM‑index and streaming only the required segments on demand, allowing a single node equipped with ≤64 GB of RAM to align a full human genome (~3 Gb).
Performance evaluation used a realistic workload of 2 TB of 150‑bp paired‑end reads (30 × coverage). In a baseline configuration, BWA‑MEM on a 32‑core CPU required roughly 3.2 hours to complete the alignment. MUSIC, deployed on a node with 16 GPUs and 2 FPGAs, finished the same task in under 15 minutes, achieving a speed‑up factor of more than 12×. Energy measurements showed a reduction of over 70 % in power consumption, indicating a markedly improved performance‑per‑watt ratio.
Importantly, MUSIC preserves full compatibility with existing BWA command‑line interfaces, allowing users to switch to the hybrid mode without modifying pipelines. The authors discuss future extensions, including containerized deployment for cloud‑based auto‑scaling and the adaptation of the hybrid acceleration strategy to other popular aligners such as Bowtie2 and Minimap2.
In summary, MUSIC demonstrates that a carefully engineered hybrid CPU‑GPU‑FPGA solution can dramatically accelerate Burrows‑Wheeler based short‑read alignment, reduce memory footprints, and lower energy costs, thereby alleviating a critical bottleneck in contemporary genomics workflows and enabling researchers to process ever‑larger sequencing datasets more efficiently.
Comments & Academic Discussion
Loading comments...
Leave a Comment