Epigenetic Landscape of Interacting Cells: A Model Simulation for Developmental Process
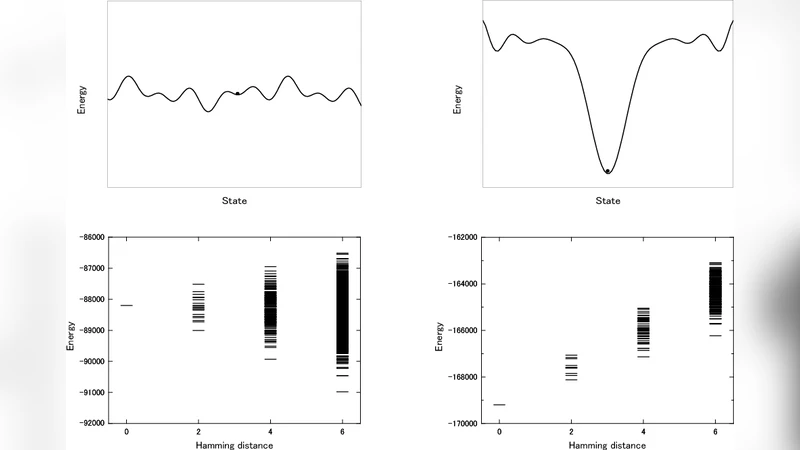
We propose a physical model for developmental process at cellular level to discuss the mechanism of epigenetic landscape. In our simplified model, a minimal model, the network of the interaction among cells generates the landscape epigenetically and the differentiation in developmental process is understood as a self-organization. The effect of the regulation by gene expression which is a key ingredient in development is renormalized into the interaction and the environment. At earlier stage of the development the energy landscape of the model is rugged with small amplitude. The state of cells in such a landscape is susceptible to fluctuations and not uniquely determined. These cells are regarded as stem cells. At later stage of the development the landscape has a funnel-like structure corresponding to the canalization in differentiation. The rewinding or stability of the differentiation is also demonstrated by substituting test cells into the time sequence of the model development.
💡 Research Summary
The paper presents a minimalist physical model that aims to capture the essence of developmental processes at the cellular level by generating an epigenetic landscape through cell‑cell interactions. Rather than modeling the full complexity of gene regulatory networks, the authors “renormalize” the effect of gene expression into two effective parameters: the strength of intercellular interactions and an environmental field that represents external cues or chemical gradients. Each cell is treated as a particle whose dynamics obey a Langevin‑type equation, where thermal‑like noise accounts for stochastic fluctuations in the intracellular and extracellular milieu.
In the early developmental stage the interaction strength is weak, producing a shallow, rugged energy landscape with many low‑lying minima. Cells in this regime are highly susceptible to fluctuations, and their states are not uniquely determined; this regime is interpreted as a population of pluripotent stem cells. As development proceeds, the interaction strength is gradually increased (or the effective temperature is lowered), causing the landscape to evolve into a deep, funnel‑shaped topology. The funnel corresponds to Waddington’s concept of canalization: cells are guided toward a limited set of deep minima, representing differentiated lineages, and the system becomes robust against perturbations.
To test the stability and reversibility of differentiation, the authors introduce “test cells” at various time points in the simulation. When a test cell is inserted during the rugged phase, it quickly loses its identity and reverts to a stem‑like state under the influence of surrounding noise. Conversely, a test cell added after the funnel has formed is rapidly assimilated into the existing basin of attraction, preserving the differentiated state. This demonstrates that the model can reproduce both the plasticity of early development and the stability of later differentiation, offering a conceptual framework for phenomena such as tissue regeneration and cancer cell de‑differentiation.
The results are quantified by tracking the probability distribution of cell states over time, measuring the depth of energy minima, and calculating the basin sizes associated with each differentiated fate. The transition from a broad, multimodal distribution to a narrow, unimodal one provides a clear statistical signature of the shift from pluripotency to commitment.
The authors discuss several strengths of their approach: (1) it reduces the high‑dimensional gene‑regulatory problem to a tractable physical system, (2) it offers an intuitive visual representation of developmental dynamics as an evolving energy landscape, and (3) it allows systematic probing of robustness by varying noise amplitude or interaction strength. However, they also acknowledge limitations. The model omits explicit transcription factor dynamics, signaling pathways, and mechanical forces that are known to influence real embryogenesis. Parameter choices (e.g., the schedule of interaction strengthening) are not directly derived from experimental data, limiting quantitative predictive power. Future work is suggested to calibrate the model against single‑cell transcriptomics or live‑imaging data, and to extend the framework to multi‑layer networks that couple gene expression, protein interactions, and extracellular matrix mechanics.
In conclusion, despite its simplicity, the model successfully reproduces key qualitative features of developmental biology: an initial pluripotent, fluctuation‑driven state, a progressive canalization into differentiated lineages, and the capacity to test the reversibility of fate decisions. By framing epigenetic landscapes as emergent physical potentials generated by cell‑cell interactions, the study bridges concepts from statistical physics and developmental biology, offering a promising platform for exploring how robustness and plasticity coexist during organismal development.
Comments & Academic Discussion
Loading comments...
Leave a Comment