Extracting tag hierarchies
Tagging items with descriptive annotations or keywords is a very natural way to compress and highlight information about the properties of the given entity. Over the years several methods have been proposed for extracting a hierarchy between the tags for systems with a “flat”, egalitarian organization of the tags, which is very common when the tags correspond to free words given by numerous independent people. Here we present a complete framework for automated tag hierarchy extraction based on tag occurrence statistics. Along with proposing new algorithms, we are also introducing different quality measures enabling the detailed comparison of competing approaches from different aspects. Furthermore, we set up a synthetic, computer generated benchmark providing a versatile tool for testing, with a couple of tunable parameters capable of generating a wide range of test beds. Beside the computer generated input we also use real data in our studies, including a biological example with a pre-defined hierarchy between the tags. The encouraging similarity between the pre-defined and reconstructed hierarchy, as well as the seemingly meaningful hierarchies obtained for other real systems indicate that tag hierarchy extraction is a very promising direction for further research with a great potential for practical applications.
💡 Research Summary
The paper addresses the problem of uncovering latent hierarchical relationships among tags in folksonomies—systems where users freely assign keywords to items, resulting in a flat, egalitarian tag space. Recognizing that many applications (search refinement, recommendation, automatic categorisation) would benefit from a structured taxonomy, the authors propose a comprehensive framework that includes (1) two novel hierarchy‑extraction algorithms, (2) a suite of quantitative quality measures, and (3) a synthetic benchmark for systematic testing.
Algorithm A builds a directed weighted co‑occurrence network where each pair of tags is represented by two opposite links weighted by the number of joint occurrences. For each tag, incoming links weaker than a fraction ω of the strongest incoming link are pruned (ω≈0.4 works well). The remaining in‑neighbors are evaluated using a z‑score that compares observed co‑occurrences to the expectation under random tagging. The neighbor with the highest z‑score is normally selected as the direct ancestor, but if that neighbor already retains a link to the current tag (indicating a sibling relationship), the algorithm scans down the z‑score list until it finds a suitable parent. A global root is chosen as the node with maximal entropy of its incoming‑weight distribution, and local roots are attached to the most frequently co‑occurring partner in a different subtree. The method runs in O(Q)+O(M log M) time, where Q is the number of objects and M the number of co‑occurrence links.
Algorithm B treats the co‑occurrence network as undirected, retaining a single weighted edge per tag pair. Links with a z‑score below 10 are removed, except when one tag appears on more than half of the objects of the other (these links are kept). Eigenvector centrality is then computed on the pruned network, and tags are processed from lowest to highest centrality. A tag’s parent must have higher centrality; if several candidates satisfy this, the algorithm aggregates z‑scores between the candidate and all already‑assigned descendants of the tag, selecting the candidate with the highest aggregate. This bottom‑up construction guarantees an acyclic hierarchy and runs in O(Q)+O(N log N) time (N = number of distinct tags).
To evaluate reconstructed hierarchies, the authors introduce several normalized metrics:
- r_E: proportion of exactly matching edges,
- r_A: proportion of edges that connect any ancestor‑descendant pair (including grandparents),
- r_U: proportion of edges linking unrelated tags,
- r_I: proportion of inverted edges (wrong direction),
- r_M: proportion of missing edges when the reconstructed graph is sparse.
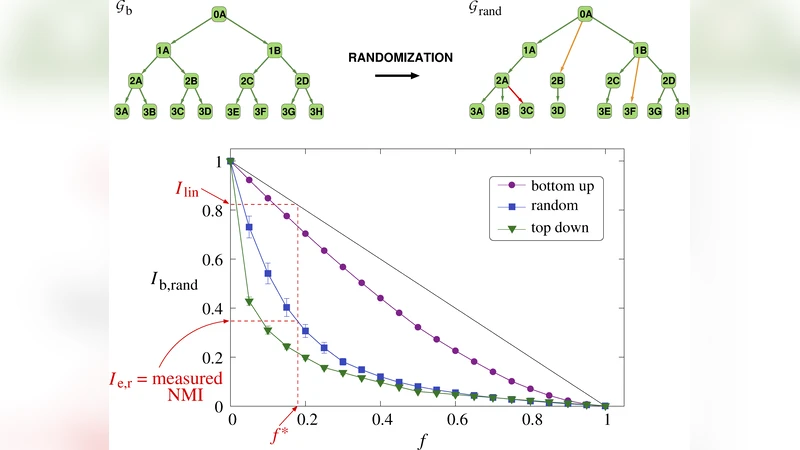
All are normalized by max(N‑1, M_r) to avoid inflation in sparse cases. Additionally, they compute normalized mutual information (NMI) between the true DAG and the reconstructed DAG, providing a global structural similarity score.
The synthetic benchmark generates virtual objects and tags according to a pre‑specified ground‑truth hierarchy. Tag frequencies follow a power‑law distribution, and the probability that a tag is assigned to an object depends on its depth in the hierarchy and a tunable noise parameter. By varying these parameters the authors test algorithm robustness under different sparsity, noise, and tag‑popularity conditions.
Empirical validation uses three real‑world datasets: (1) protein function annotations from the Gene Ontology, where the true hierarchy is known; (2) Flickr photo tags; and (3) IMDb movie tags. On the protein data, Algorithm A achieves the highest r_E and NMI, reflecting its strength when the underlying hierarchy is relatively shallow and well‑defined. On the synthetic benchmark and the two social‑media datasets, Algorithm B consistently outperforms A, producing more coherent and semantically plausible trees, as judged both by quantitative metrics and visual inspection.
Overall, the study delivers a full pipeline—from data generation and algorithmic extraction to multi‑faceted evaluation—for tag hierarchy reconstruction. It demonstrates that simple co‑occurrence statistics, when combined with careful statistical filtering (z‑score thresholds) and network‑theoretic centrality measures, can reliably infer meaningful taxonomies from flat tag collections. Future work suggested includes handling dynamic tag streams, integrating synonym/antonym relations, and scaling the methods to massive, real‑time systems.
Comments & Academic Discussion
Loading comments...
Leave a Comment