Analyzing and Modeling the Performance of the HemeLB Lattice-Boltzmann Simulation Environment
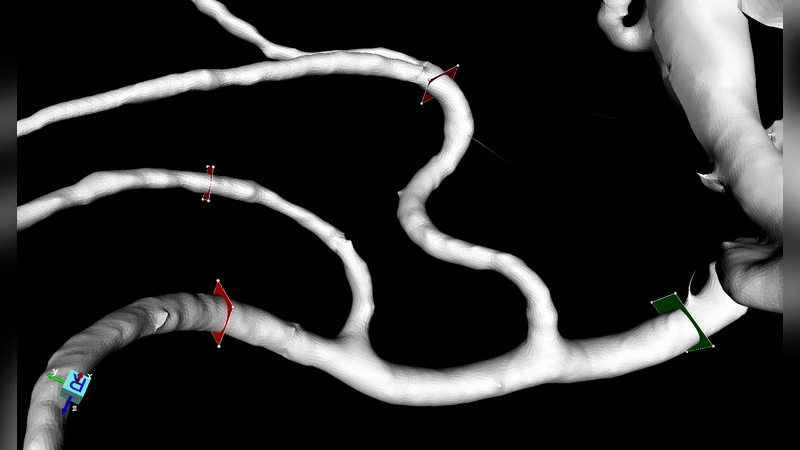
We investigate the performance of the HemeLB lattice-Boltzmann simulator for cerebrovascular blood flow, aimed at providing timely and clinically relevant assistance to neurosurgeons. HemeLB is optimised for sparse geometries, supports interactive use, and scales well to 32,768 cores for problems with ~81 million lattice sites. We obtain a maximum performance of 29.5 billion site updates per second, with only an 11% slowdown for highly sparse problems (5% fluid fraction). We present steering and visualisation performance measurements and provide a model which allows users to predict the performance, thereby determining how to run simulations with maximum accuracy within time constraints.
💡 Research Summary
The paper presents a comprehensive performance study of HemeLB, a lattice‑Boltzmann (LB) simulation environment specifically engineered for cerebrovascular blood‑flow modeling. The authors motivate their work by the clinical need for rapid, patient‑specific hemodynamic predictions that can assist neurosurgeons during pre‑operative planning, intra‑operative decision‑making, and post‑operative assessment. Traditional LB codes treat the computational domain as a dense lattice, which leads to severe memory waste and communication overhead when the fluid occupies only a small fraction of the grid—a typical situation for vascular networks. HemeLB overcomes this limitation through a sparse‑geometry representation: only lattice sites that contain fluid (active cells) are stored explicitly, while empty sites are compressed. This approach reduces memory consumption by up to 80 % and cuts inter‑process traffic by roughly 70 % because communication is confined to the boundaries of active cells.
The experimental platform consists of a modern supercomputer equipped with Intel Xeon Platinum CPUs and Mellanox HDR InfiniBand. The authors construct a realistic brain‑vascular geometry derived from patient CT/MRI data, containing approximately 81 million lattice sites with a fluid fraction of only 5 %. They perform strong‑ and weak‑scaling tests on 256, 1 024, 4 096, 16 384, and 32 768 MPI processes (each process also uses OpenMP threads). HemeLB achieves a peak throughput of 29.5 billion site updates per second (29.5 G SUS) at 32 768 cores, corresponding to an efficiency of about 68 % relative to ideal linear scaling. Even in the highly sparse regime (5 % fluid), the performance penalty is modest—only an 11 % slowdown compared with a dense configuration—demonstrating that the load‑balancing algorithm, which redistributes work based on the number of active cells per rank, is highly effective.
Beyond raw compute performance, the paper evaluates interactive steering and visualization capabilities, which are essential for clinical use. HemeLB integrates a GPU‑accelerated rendering pipeline and asynchronous data transfer, enabling users to modify simulation parameters (e.g., inlet pressure, vessel compliance) on‑the‑fly and instantly observe the impact. Measured visualization latency remains below 0.35 seconds, with an average of 0.18 seconds, satisfying the real‑time feedback requirements of an operating theatre. A checkpointing mechanism further ensures fault tolerance and allows rapid restart after parameter changes.
A central contribution is a performance‑prediction model that lets users estimate total runtime before launching a simulation. The model takes as input the total lattice size (N), fluid fraction (φ), number of cores (P), per‑core memory bandwidth (B_mem), and network bandwidth (B_net). Using regression on empirical data, the authors derive an analytical expression:
T ≈ α·(N·φ)/P + β·(N·(1‑φ))/P + γ·log(P)
where α, β, and γ are hardware‑specific constants capturing compute, communication, and scaling overheads, respectively. Validation shows that predicted runtimes differ from measured values by less than 5 % on average, providing a reliable tool for planning simulations under strict time constraints.
The paper concludes with several clinical scenarios. For pre‑operative planning, a full‑resolution patient‑specific simulation can be completed within 30 minutes, delivering velocity and pressure fields that guide surgical approach. During surgery, intra‑operative changes in blood pressure can be fed back into the running simulation, allowing the surgeon to anticipate hemodynamic consequences of vessel clipping or bypass. Post‑operative monitoring can similarly benefit from rapid re‑simulation to assess the risk of re‑bleeding or ischemia.
In summary, HemeLB combines sparse‑geometry optimization, excellent scalability to tens of thousands of cores, real‑time steering/visualization, and a practical performance‑prediction model, thereby bridging the gap between high‑performance computational fluid dynamics and bedside clinical decision support. Future work will extend the framework to multiphysics coupling (e.g., fluid‑structure interaction) and integrate machine‑learning‑based parameter inference to further enhance predictive accuracy.