MAGNA: Maximizing Accuracy in Global Network Alignment
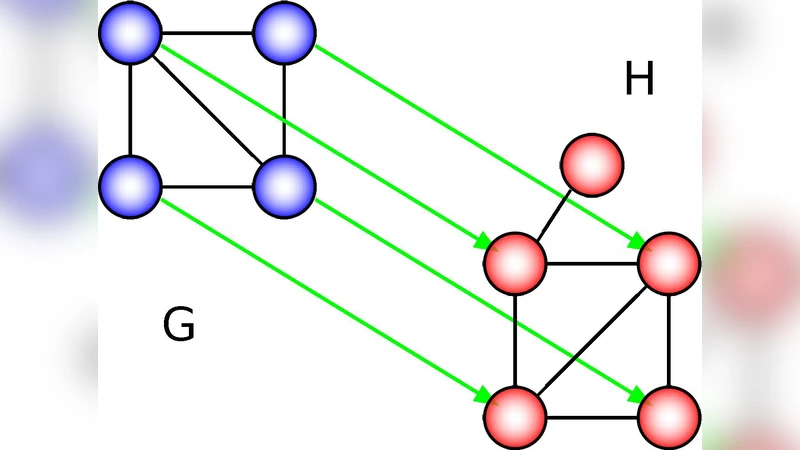
Biological network alignment aims to identify similar regions between networks of different species. Existing methods compute node “similarities” to rapidly identify from possible alignments the “high-scoring” alignments with respect to the overall node similarity. However, the accuracy of the alignments is then evaluated with some other measure that is different than the node similarity used to construct the alignments. Typically, one measures the amount of conserved edges. Thus, the existing methods align similar nodes between networks hoping to conserve many edges (after the alignment is constructed!). Instead, we introduce MAGNA to directly “optimize” edge conservation while the alignment is constructed. MAGNA uses a genetic algorithm and our novel function for crossover of two “parent” alignments into a superior “child” alignment to simulate a “population” of alignments that “evolves” over time; the “fittest” alignments survive and proceed to the next “generation”, until the alignment accuracy cannot be optimized further. While we optimize our new and superior measure of the amount of conserved edges, MAGNA can optimize any alignment accuracy measure. In systematic evaluations against existing state-of-the-art methods (IsoRank, MI-GRAAL, and GHOST), MAGNA improves alignment accuracy of all methods.
💡 Research Summary
The paper introduces MAGNA (Maximizing Accuracy in Global Network Alignment), a novel framework for global network alignment (GNA) that directly optimizes edge‑conservation during the alignment construction process. Traditional GNA methods follow a two‑step pipeline: first they compute similarity scores for all possible node pairs (using spectral methods such as IsoRank, graphlet‑based scores as in the GRAAL family, or spectral signatures as in GHOST), and then they search for high‑scoring alignments based on those node similarity scores. The quality of the resulting alignment, however, is typically evaluated with a different metric—most commonly the amount of conserved edges (edge correctness, EC) or related measures such as induced conserved structure (ICS) and symmetric substructure score (S³). This mismatch between the objective used to build the alignment and the objective used to evaluate it can lead to sub‑optimal results.
MAGNA addresses this fundamental inconsistency by employing a genetic algorithm (GA) whose fitness function is precisely the edge‑conservation measure one wishes to maximize. Alignments are represented as permutations (bijections) between the node sets of the two networks; when the networks differ in size, dummy zero‑degree nodes are added to the smaller network so that a total injective mapping becomes a permutation of size n. The core of MAGNA is a novel crossover operator defined on the permutation space. Two parent permutations σ and τ are considered adjacent if they differ by a single transposition. By constructing a graph Γₙ whose vertices are all permutations and whose edges connect adjacent permutations, the crossover σ⊗τ is defined as the midpoint on a shortest path between σ and τ in Γₙ. Theoretical analysis shows that, for large n, the child permutation shares roughly half of its node pairs with each parent, providing a balanced inheritance of structural features.
The GA proceeds as follows. An initial population of alignments is generated in four ways: (1) completely random permutations, (2) random permutations mixed with an IsoRank alignment, (3) random permutations mixed with a MI‑GRAAL alignment, and (4) random permutations mixed with a GHOST alignment. Population sizes ranging from 200 to 15 000 are explored. In each generation, the top 50 % of the population (the “elite”) is retained unchanged, while the remaining slots are filled with offspring produced by the crossover operator. Parent pairs are selected via roulette‑wheel selection, which gives higher‑fitness alignments a proportionally larger chance of being chosen. The process repeats for many generations until the fitness (edge‑conservation) plateaus.
MAGNA’s fitness can be any alignment quality measure; the authors experiment with EC, ICS, and S³, and also propose a composite metric that captures the strengths of existing measures. Because the GA framework is agnostic to the specific metric, MAGNA can be adapted to optimize purely topological, purely biological (e.g., sequence similarity), or hybrid objectives.
Experimental evaluation uses a high‑confidence yeast protein‑protein interaction (PPI) network and several noisy variants where the true node mapping is known. This benchmark allows a direct comparison of alignment accuracy. MAGNA is compared against three state‑of‑the‑art GNA methods: IsoRank, MI‑GRAAL, and GHOST. Results show that, across all population sizes and initializations, MAGNA consistently achieves higher EC, higher ICS, and higher S³ than the baseline methods. When an existing method’s alignment is included in the initial population, convergence accelerates and final quality improves further, demonstrating that MAGNA can both refine existing alignments and generate superior ones from scratch.
The paper’s contributions are fivefold: (1) redefining the GNA objective to directly maximize edge‑conservation during alignment construction, (2) introducing the first permutation‑based crossover operator for network alignment, (3) being the first to apply a genetic algorithm to PPI GNA, (4) proposing a new composite alignment quality metric that outperforms individual measures, and (5) providing extensive empirical validation across multiple population configurations and baseline methods. By aligning the optimization target with the evaluation metric, MAGNA establishes a new design principle for future network alignment algorithms. Moreover, its flexible fitness function opens the door to multi‑objective alignments that simultaneously consider topological and biological information, potentially benefiting applications such as cross‑species functional annotation transfer, disease‑gene discovery, and the integration of heterogeneous biological networks.
Comments & Academic Discussion
Loading comments...
Leave a Comment