Nonparametric Multi-group Membership Model for Dynamic Networks
Relational data-like graphs, networks, and matrices-is often dynamic, where the relational structure evolves over time. A fundamental problem in the analysis of time-varying network data is to extract a summary of the common structure and the dynamics of the underlying relations between the entities. Here we build on the intuition that changes in the network structure are driven by the dynamics at the level of groups of nodes. We propose a nonparametric multi-group membership model for dynamic networks. Our model contains three main components: We model the birth and death of individual groups with respect to the dynamics of the network structure via a distance dependent Indian Buffet Process. We capture the evolution of individual node group memberships via a Factorial Hidden Markov model. And, we explain the dynamics of the network structure by explicitly modeling the connectivity structure of groups. We demonstrate our model’s capability of identifying the dynamics of latent groups in a number of different types of network data. Experimental results show that our model provides improved predictive performance over existing dynamic network models on future network forecasting and missing link prediction.
💡 Research Summary
The paper introduces a novel Bayesian non‑parametric framework for modeling dynamic relational data such as time‑varying graphs, networks, and matrices. The authors argue that changes in network structure are best explained at the level of groups of nodes rather than at the level of individual edges. Accordingly, they propose a three‑component model: (1) a birth‑and‑death process for latent groups, (2) a factorial hidden Markov model (FHMM) for the evolution of node‑group memberships, and (3) a link function that explicitly incorporates both intra‑group and inter‑group interactions.
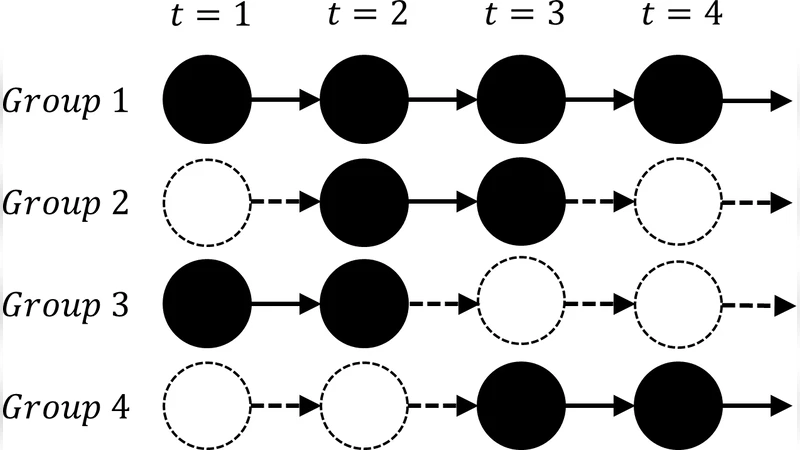
Group birth and death are modeled with a distance‑dependent Indian Buffet Process (dd‑IBP). Each discrete time step is treated as a “customer” that samples a set of active groups. At t = 1 the number of active groups follows a Poisson(λ) distribution. For t > 1, each currently active group becomes inactive with probability γ, while a Poisson(γλ) number of brand‑new groups are born. Once a group becomes inactive it never revives, which eliminates the ambiguity between a group’s disappearance and a mere change in its membership composition. The hyper‑parameter γ controls the temporal coherence of groups: γ≈1 yields long‑lived groups, γ≈0 yields a rapidly changing group landscape.
Node‑group memberships are binary variables z_{ik}^{(t)}∈{0,1}. For each group k, the membership of node i evolves according to an independent 2×2 Markov transition matrix Q_k. The two off‑diagonal entries of Q_k are denoted a_k (probability of joining) and b_k (probability of leaving) and are given Beta(α,β) priors, typically with β>α to encourage relatively stable memberships. This FHMM formulation allows a node to belong to multiple groups simultaneously, avoiding the normalization constraint of mixed‑membership models that force a trade‑off between groups.
The link generation mechanism builds on the Multiplicative Attribute Graph (MAG) idea. Each group k possesses a 2×2 affinity matrix Θ_k whose entries correspond to the four possible membership combinations between a pair of nodes (member‑member, member‑non‑member, non‑member‑member, non‑member‑non‑member). For a pair (i,j) at time t, the relevant entry Θ_k
Comments & Academic Discussion
Loading comments...
Leave a Comment