Mapping of the Influenza-A Hemagglutinin Serotypes Evolution by the ISSCOR Method
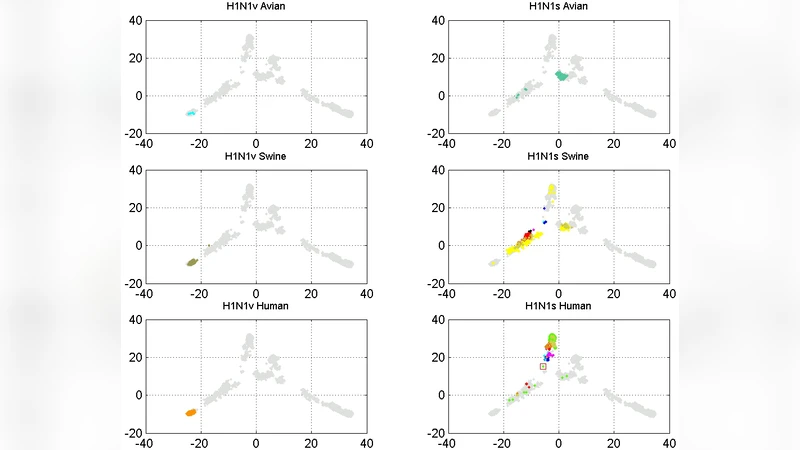
Analyses and visualizations by the ISSCOR method of influenza virus hemagglutinin genes of different A-subtypes revealed some rather striking temporal relationships between groups of individual gene subsets. Based on these findings we consider application of the ISSCOR-PCA method for analyses of large sets of homologous genes to be a worthwhile addition to a toolbox of genomics - allowing for a rapid diagnostics of trends, and ultimately even aiding an early warning of newly emerging epidemiological threats.
💡 Research Summary
**
The paper introduces a novel computational framework that combines the ISSCOR (Intragenic Stochastic Synonymous Codon Occurrence Replacement) method with principal component analysis (PCA) to investigate the evolutionary dynamics of influenza A virus hemagglutinin (HA) genes. ISSCOR generates ensembles of artificial nucleotide sequences that retain the exact amino‑acid translation of an original HA gene while randomly shuffling synonymous codons according to their usage frequencies. By comparing the observed frequencies of codon‑pair patterns (including various spacer lengths λ) in the original sequence with those in a large set of shuffled replicas, the authors compute standardized deviates (Txx) that quantify how far the native codon order deviates from a random expectation.
Using 9,131 full‑length HA sequences collected from the NCBI influenza resource (covering subtypes H1N1, H1N1v (2009‑2020 pandemic), H1N2, H2Nx, H3N2, H5N1, H7Nx and hosts such as avian, human, swine, and ferret), the authors calculate deviates for λ ranging from 0 to 16, yielding a 2,448‑dimensional feature vector for each gene. PCA on this matrix shows that the first two components explain 45.4 % and 18.3 % of the total variance, respectively, allowing the entire dataset to be visualized in a two‑dimensional space.
The resulting maps reveal clear, host‑ and subtype‑specific clusters. H3N2 sequences form a well‑ordered trajectory that mirrors the classic avian → swine → human transmission route. In contrast, the 2009 pandemic H1N1 isolates display a reversed pattern: a few avian isolates from late 2009 (Canadian turkey samples) cluster together with human and swine strains, suggesting a direct avian‑to‑human jump that bypasses the swine intermediate. Other subtypes (H1N1, H5N1, H1N2, H2Nx, H7Nx) exhibit a more tangled network of points, indicating multiple possible evolutionary pathways and frequent intermixing among hosts.
When all points are plotted together, they outline an approximate triangle: the oldest sequences (e.g., the 1918 H1N1) occupy the upper vertex, the most recent H3N2 strains lie at the lower right, and the newest 2009 pandemic H1N1 strains sit at the lower left. This geometric arrangement provides an intuitive visual chronology of HA evolution over the past century.
Methodologically, the authors validate ISSCOR‑PCA results against traditional phylogenetic approaches (Neighbor‑Joining, QPF, and MUSCLE‑based multiple alignments) and find substantial concordance, demonstrating that a non‑alignment, order‑based descriptor can capture biologically meaningful relationships. They discuss several limitations: ISSCOR ignores selective pressures related to RNA secondary structure, translation efficiency, and immune evasion; higher‑order λ values suffer from sparsity; PCA captures only linear variance; and the underlying sequence database may be biased by geographic and temporal sampling.
Despite these caveats, the study showcases ISSCOR‑PCA as a powerful, scalable tool for extracting subtle codon‑order signals from large genomic collections. The authors propose that the approach could be extended to other rapidly evolving pathogens (e.g., coronaviruses) and integrated into early‑warning pipelines for emerging viral threats, complementing existing phylogenetic and epidemiological surveillance methods.
Comments & Academic Discussion
Loading comments...
Leave a Comment